Model Internals
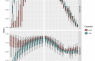
Each PhenDB model is trained on sets of bacterial ENOGs (orthologous groups from EggNOG 4.5), which have or have not been identified in the training genomes. Each ENOG is given a weight, with the magnitude of the weight being the importance of that ENOG for the final prediction. The sign of the weight indicates whether the presence (positive weight) or absence (negative weight) of this ENOG is indicative of the trait.
This table lists the 250 highest-ranking ENOGs of this model.
| rank in model | enog name | enog description | weight in model |
|---|---|---|---|
| 1 | COG1720 | tRNA m6t6A37 methyltransferase activity | 0.023790 |
| 2 | COG2842 | Transposition protein | 0.022884 |
| 3 | COG2046 | Belongs to the sulfate adenylyltransferase family | 0.022296 |
| 4 | COG1348 | The key enzymatic reactions in nitrogen fixation are catalyzed by the nitrogenase complex which has 2 components the iron protein and the molybdenum-iron protein | 0.021227 |
| 5 | COG2710 | Component of the dark-operative protochlorophyllide reductase (DPOR) that uses Mg-ATP and reduced ferredoxin to reduce ring D of protochlorophyllide (Pchlide) to form chlorophyllide a (Chlide). This reaction is light-independent. The NB-protein | 0.021227 |
| 6 | COG1200 | Critical role in recombination and DNA repair. Helps process Holliday junction intermediates to mature products by catalyzing branch migration. Has a DNA unwinding activity characteristic of a DNA helicase with a 3'- to 5'- polarity. Unwinds branched duplex DNA (Y-DNA) | -0.021019 |
| 7 | COG1940 | carbohydrate kinase activity | -0.021007 |
| 8 | COG1053 | succinate dehydrogenase | 0.020945 |
| 9 | 339S5 | Putative regulatory protein | 0.020283 |
| 10 | COG1900 | Homocysteine biosynthesis enzyme, sulfur-incorporation | 0.020125 |
| 11 | COG1035 | Coenzyme F420 hydrogenase/dehydrogenase, beta subunit C terminus | 0.019817 |
| 12 | COG3505 | Type IV secretory pathway VirD4 | 0.019766 |
| 13 | COG4087 | Haloacid dehalogenase domain protein hydrolase | 0.019499 |
| 14 | COG1916 | peptidoglycan binding | 0.019342 |
| 15 | COG5561 | CGGC domain | 0.019024 |
| 16 | COG0578 | Belongs to the FAD-dependent glycerol-3-phosphate dehydrogenase family | 0.018809 |
| 17 | COG1489 | DNA binding | 0.018714 |
| 18 | COG2122 | PFAM ApbE family | 0.018622 |
| 19 | COG2873 | o-acetylhomoserine | -0.018613 |
| 20 | COG0857 | belongs to the CobB CobQ family | 0.018407 |
| 21 | 337H4 | 0.018334 | |
| 22 | COG0785 | Cytochrome C biogenesis protein | -0.018169 |
| 23 | COG0555 | PFAM binding-protein-dependent transport systems inner membrane component | -0.018166 |
| 24 | 32TNY | Domain of unknown function (DUF1992) | 0.018149 |
| 25 | COG2104 | thiamine diphosphate biosynthetic process | -0.018049 |
| 26 | COG3833 | ABC-type maltose transport systems, permease component | -0.017973 |
| 27 | COG3705 | Required for the first step of histidine biosynthesis. May allow the feedback regulation of ATP phosphoribosyltransferase activity by histidine | -0.017972 |
| 28 | COG0855 | Catalyzes the reversible transfer of the terminal phosphate of ATP to form a long-chain polyphosphate (polyP) | -0.017964 |
| 29 | COG0276 | Catalyzes the ferrous insertion into protoporphyrin IX | -0.017666 |
| 30 | COG0230 | Belongs to the bacterial ribosomal protein bL34 family | -0.017577 |
| 31 | COG2920 | part of a sulfur-relay system | 0.017344 |
| 32 | COG4536 | flavin adenine dinucleotide binding | -0.017300 |
| 33 | COG2043 | Uncharacterised ArCR, COG2043 | 0.017214 |
| 34 | COG3933 | PFAM sigma-54 factor interaction domain-containing protein | -0.017150 |
| 35 | COG2145 | Catalyzes the phosphorylation of the hydroxyl group of 4-methyl-5-beta-hydroxyethylthiazole (THZ) | 0.017034 |
| 36 | COG0248 | Ppx GppA phosphatase | -0.016675 |
| 37 | COG2998 | abc-type tungstate transport system, permease component | 0.016533 |
| 38 | COG3127 | ABC-type transport system involved in lysophospholipase L1, biosynthesis, permease component | -0.016502 |
| 39 | COG3418 | PFAM FlgN family protein | -0.016492 |
| 40 | COG2221 | Nitrite and sulphite reductase 4Fe-4S | 0.016469 |
| 41 | COG1578 | Protein of unknown function DUF89 | 0.016393 |
| 42 | COG1359 | (4S)-4-hydroxy-5-phosphonooxypentane-2,3-dione isomerase activity | 0.016337 |
| 43 | 32RXU | RsbT co-antagonist protein rsbRD N-terminal domain | 0.016311 |
| 44 | 32SEM | 0.016311 | |
| 45 | COG3857 | The heterodimer acts as both an ATP-dependent DNA helicase and an ATP-dependent, dual-direction single-stranded exonuclease. Recognizes the chi site generating a DNA molecule suitable for the initiation of homologous recombination | -0.016275 |
| 46 | 32S9W | Protein of unknown function (DUF3189) | -0.016269 |
| 47 | COG1105 | Belongs to the carbohydrate kinase PfkB family | -0.016082 |
| 48 | COG0534 | Mate efflux family protein | -0.016080 |
| 49 | COG4951 | DNA primase small subunit | 0.015997 |
| 50 | COG2232 | ATP-grasp domain | 0.015853 |
| 51 | COG2362 | D-Aminopeptidase | -0.015838 |
| 52 | COG0308 | aminopeptidase N | -0.015762 |
| 53 | COG2972 | Histidine kinase | -0.015635 |
| 54 | COG3808 | pump that utilizes the energy of pyrophosphate hydrolysis as the driving force for | -0.015627 |
| 55 | COG1906 | membrane | 0.015620 |
| 56 | COG0475 | glutathione-regulated potassium exporter activity | 0.015489 |
| 57 | COG0661 | Is probably a protein kinase regulator of UbiI activity which is involved in aerobic coenzyme Q (ubiquinone) biosynthesis | -0.015448 |
| 58 | COG1908 | Methyl-viologen-reducing hydrogenase, delta subunit | 0.015428 |
| 59 | COG3271 | cysteine-type peptidase activity | 0.015368 |
| 60 | COG4129 | membrane | 0.015296 |
| 61 | 3224Y | Universal stress protein family | 0.015275 |
| 62 | COG3075 | Conversion of glycerol 3-phosphate to dihydroxyacetone. Uses fumarate or nitrate as electron acceptor | 0.015272 |
| 63 | COG2107 | Catalyzes the conversion of cyclic dehypoxanthine futalosine (cyclic DHFL) into 1,4-dihydroxy-6-naphthoate, a step in the biosynthesis of menaquinone (MK, vitamin K2) | 0.015257 |
| 64 | 32SDN | 0.015181 | |
| 65 | 32VSP | 0.015123 | |
| 66 | COG0693 | protein deglycase activity | 0.014961 |
| 67 | COG1683 | Conserved Protein | 0.014933 |
| 68 | COG1351 | Catalyzes the reductive methylation of 2'-deoxyuridine- 5'-monophosphate (dUMP) to 2'-deoxythymidine-5'-monophosphate (dTMP) while utilizing 5,10-methylenetetrahydrofolate (mTHF) as the methyl donor, and NAD(P)H and FADH(2) as the reductant | -0.014925 |
| 69 | COG0646 | methionine synthase | -0.014828 |
| 70 | COG2189 | Belongs to the N(4) N(6)-methyltransferase family | -0.014808 |
| 71 | COG1136 | (ABC) transporter | -0.014785 |
| 72 | 33AT3 | Sulfurtransferase TusA | 0.014764 |
| 73 | COG0031 | Belongs to the cysteine synthase cystathionine beta- synthase family | -0.014730 |
| 74 | COG0220 | tRNA (guanine-N7-)-methyltransferase activity | -0.014706 |
| 75 | COG2864 | formate dehydrogenase | -0.014683 |
| 76 | COG1741 | Belongs to the pirin family | -0.014645 |
| 77 | COG1803 | Methylglyoxal synthase | -0.014617 |
| 78 | COG0314 | molybdopterin synthase activity | 0.014553 |
| 79 | COG4812 | Cobalamin adenosyltransferase | 0.014552 |
| 80 | COG3655 | Transcriptional regulator | 0.014518 |
| 81 | COG1573 | uracil-dna glycosylase | -0.014497 |
| 82 | COG1893 | Catalyzes the NADPH-dependent reduction of ketopantoate into pantoic acid | -0.014393 |
| 83 | COG0257 | Belongs to the bacterial ribosomal protein bL36 family | -0.014299 |
| 84 | COG1737 | Transcriptional regulator | -0.014222 |
| 85 | 32TK9 | 0.014167 | |
| 86 | COG3173 | Aminoglycoside phosphotransferase | 0.014075 |
| 87 | COG5350 | Protein tyrosine phosphatase | 0.014059 |
| 88 | COG2956 | Modulates cellular lipopolysaccharide (LPS) levels by regulating LpxC, which is involved in lipid A biosynthesis. May act by modulating the proteolytic activity of FtsH towards LpxC. May also coordinate assembly of proteins involved in LPS synthesis at the plasma membrane | -0.014047 |
| 89 | COG4666 | Tripartite ATP-independent periplasmic transporter, DctM component | 0.014013 |
| 90 | 307WW | 0.013988 | |
| 91 | COG4822 | anaerobic cobalamin biosynthetic process | 0.013983 |
| 92 | COG4766 | ethanolamine catabolic process | 0.013958 |
| 93 | COG1434 | Gram-negative-bacterium-type cell wall biogenesis | -0.013954 |
| 94 | COG1323 | Belongs to the UPF0348 family | -0.013907 |
| 95 | COG0607 | Catalyzes the ATP-dependent transfer of a sulfur to tRNA to produce 4-thiouridine in position 8 of tRNAs, which functions as a near-UV photosensor. Also catalyzes the transfer of sulfur to the sulfur carrier protein ThiS, forming ThiS-thiocarboxylate. This is a step in the synthesis of thiazole, in the thiamine biosynthesis pathway. The sulfur is donated as persulfide by IscS | -0.013906 |
| 96 | COG2876 | 3-deoxy-7-phosphoheptulonate synthase activity | -0.013806 |
| 97 | 33P2T | 0.013803 | |
| 98 | 31CNF | 0.013800 | |
| 99 | 33IU2 | 0.013726 | |
| 100 | COG4917 | ethanolamine utilization protein | 0.013647 |
| 101 | COG4637 | Psort location Cytoplasmic, score | -0.013625 |
| 102 | COG0626 | cystathionine gamma-synthase activity | -0.013613 |
| 103 | COG0281 | malic enzyme | -0.013591 |
| 104 | COG2014 | Putative heavy-metal chelation | 0.013550 |
| 105 | COG1386 | Participates in chromosomal partition during cell division. May act via the formation of a condensin-like complex containing Smc and ScpA that pull DNA away from mid-cell into both cell halves | -0.013502 |
| 106 | COG4012 | pyruvate format-lyase activating enzyme | 0.013453 |
| 107 | COG0511 | ligase activity, forming carbon-carbon bonds | -0.013452 |
| 108 | COG0070 | glutamate synthase | -0.013382 |
| 109 | COG3192 | ethanolamine utilization protein | 0.013313 |
| 110 | COG4174 | ABC transporter (Permease) | 0.013307 |
| 111 | COG0209 | Provides the precursors necessary for DNA synthesis. Catalyzes the biosynthesis of deoxyribonucleotides from the corresponding ribonucleotides | -0.013261 |
| 112 | COG0145 | ligase activity | 0.013251 |
| 113 | 31H63 | 0.013216 | |
| 114 | 32SHU | Domain of unknown function (DUF2760) | 0.013215 |
| 115 | COG0337 | Catalyzes the conversion of 3-deoxy-D-arabino- heptulosonate 7-phosphate (DAHP) to dehydroquinate (DHQ) | -0.013203 |
| 116 | COG1691 | (AIR) carboxylase | 0.013187 |
| 117 | COG2895 | sulfate adenylyltransferase (ATP) activity | -0.013172 |
| 118 | COG0471 | metal ion transport | 0.013113 |
| 119 | 2ZK16 | Histidine Phosphotransfer domain | 0.013094 |
| 120 | 2ZXZV | Outer membrane efflux protein | -0.013087 |
| 121 | COG2193 | ferroxidase activity | -0.013075 |
| 122 | 32CXY | phospholipase C | 0.013075 |
| 123 | 2ZHJE | Cyclic nucleotide-binding domain | -0.013065 |
| 124 | COG4820 | ethanolamine utilization protein (EutJ) | 0.013047 |
| 125 | COG2868 | ribosomal protein | -0.013035 |
| 126 | 2Z9T8 | Domain of unknown function (DUF4405) | 0.013027 |
| 127 | 32U1K | 0.012993 | |
| 128 | 32TGZ | 0.012992 | |
| 129 | 33CDW | 0.012992 | |
| 130 | COG1350 | The beta subunit is responsible for the synthesis of L- tryptophan from indole and L-serine | -0.012973 |
| 131 | COG0118 | IGPS catalyzes the conversion of PRFAR and glutamine to IGP, AICAR and glutamate. The HisH subunit provides the glutamine amidotransferase activity that produces the ammonia necessary to HisF for the synthesis of IGP and AICAR | -0.012960 |
| 132 | 2ZVPG | 0.012939 | |
| 133 | 30F9Z | 0.012927 | |
| 134 | COG1188 | Ribosome-associated heat shock protein implicated in the recycling of the 50S subunit, (S4 paralog)) | -0.012890 |
| 135 | COG0302 | gtp cyclohydrolase | -0.012864 |
| 136 | 32ZJH | Domain of unknown function (DUF4363) | 0.012858 |
| 137 | COG1641 | Involved in the biosynthesis of a nickel-pincer cofactor ((SCS)Ni(II) pincer complex). Binds Ni(2 ), and functions in nickel delivery to pyridinium-3,5-bisthiocarboxylic acid mononucleotide (P2TMN), to form the mature cofactor. Is thus probably required for the activation of nickel-pincer cofactor- dependent enzymes | 0.012854 |
| 138 | COG3426 | butyrate kinase activity | 0.012832 |
| 139 | 33ASE | 0.012831 | |
| 140 | 32SN5 | 0.012826 | |
| 141 | 2ZCJT | 0.012815 | |
| 142 | COG1427 | Catalyzes the dehydration of chorismate into 3- (1- carboxyvinyl)oxy benzoate, a step in the biosynthesis of menaquinone (MK, vitamin K2) | 0.012813 |
| 143 | COG4745 | 4-amino-4-deoxy-L-arabinose transferase activity | -0.012769 |
| 144 | COG2132 | Multicopper oxidase | -0.012757 |
| 145 | COG1058 | Belongs to the CinA family | -0.012741 |
| 146 | COG4810 | utilization protein | 0.012702 |
| 147 | 33191 | PFAM Dissimilatory sulfite reductase D | 0.012665 |
| 148 | COG3836 | Belongs to the HpcH HpaI aldolase family | -0.012628 |
| 149 | COG0761 | Catalyzes the conversion of 1-hydroxy-2-methyl-2-(E)- butenyl 4-diphosphate (HMBPP) into a mixture of isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). Acts in the terminal step of the DOXP MEP pathway for isoprenoid precursor biosynthesis | -0.012586 |
| 150 | COG1150 | heterodisulfide reductase subunit C | 0.012527 |
| 151 | 2Z9E1 | PFAM TnsA endonuclease | 0.012523 |
| 152 | COG3108 | Peptidase M15 | 0.012481 |
| 153 | COG4108 | Increases the formation of ribosomal termination complexes and stimulates activities of RF-1 and RF-2. It binds guanine nucleotides and has strong preference for UGA stop codons. It may interact directly with the ribosome. The stimulation of RF- 1 and RF-2 is significantly reduced by GTP and GDP, but not by GMP | 0.012479 |
| 154 | 33EUU | 0.012441 | |
| 155 | COG1301 | dicarboxylic acid transport | -0.012431 |
| 156 | COG3688 | RNA-binding protein containing a PIN domain | 0.012423 |
| 157 | COG2048 | Heterodisulfide reductase, subunit B | 0.012415 |
| 158 | 3482A | -0.012406 | |
| 159 | COG1512 | COG1512 Beta-propeller domains of methanol dehydrogenase type | -0.012383 |
| 160 | COG0757 | Catalyzes a trans-dehydration via an enolate intermediate | -0.012365 |
| 161 | COG3328 | transposase activity | -0.012321 |
| 162 | COG2151 | metal-sulfur cluster biosynthetic enzyme | -0.012266 |
| 163 | COG1928 | C-terminal four TMM region of protein-O-mannosyltransferase | 0.012258 |
| 164 | COG5266 | PFAM Nickel transport complex, NikM subunit, transmembrane | 0.012249 |
| 165 | 310ZN | Protein of unknown function (DUF3795) | 0.012194 |
| 166 | 33MTN | 0.012171 | |
| 167 | COG1612 | heme a metabolic process | -0.012171 |
| 168 | COG4665 | TRAP-type mannitol chloroaromatic compound transport system small permease component | 0.012167 |
| 169 | COG4936 | Histidine kinase | 0.012137 |
| 170 | COG1825 | This is one of the proteins that binds to the 5S RNA in the ribosome where it forms part of the central protuberance | -0.012133 |
| 171 | COG4619 | ABC transporter | 0.012114 |
| 172 | COG4548 | von Willebrand factor, type A | 0.012088 |
| 173 | COG3454 | Phosphonate metabolism | 0.012060 |
| 174 | COG4799 | Acetyl-CoA carboxylase, carboxyltransferase component subunits alpha and beta | -0.012057 |
| 175 | COG1329 | Transcriptional regulator | 0.012048 |
| 176 | COG3848 | Pyruvate phosphate dikinase | 0.012010 |
| 177 | 33JWE | 0.011991 | |
| 178 | COG2274 | protein secretion by the type I secretion system | -0.011989 |
| 179 | COG4302 | ethanolamine ammonia-lyase activity | 0.011975 |
| 180 | 33WYX | 0.011972 | |
| 181 | COG3593 | DNA synthesis involved in DNA repair | 0.011962 |
| 182 | COG1704 | LemA family | -0.011961 |
| 183 | COG1598 | PFAM Uncharacterised protein family UPF0150 | 0.011916 |
| 184 | COG4963 | Pilus assembly protein | 0.011897 |
| 185 | 33BA1 | 0.011896 | |
| 186 | COG0428 | transporter | -0.011869 |
| 187 | 30SQC | PFAM UspA | 0.011861 |
| 188 | COG1610 | YqeY-like protein | -0.011859 |
| 189 | COG1253 | flavin adenine dinucleotide binding | 0.011855 |
| 190 | COG1310 | metal-dependent protease of the Pad1 Jab1 superfamily | -0.011806 |
| 191 | COG2080 | 2 iron, 2 sulfur cluster binding | -0.011798 |
| 192 | 303FQ | TIGRFAM C_GCAxxG_C_C family protein | 0.011792 |
| 193 | COG3087 | Lytic transglycosylase with a strong preference for naked glycan strands that lack stem peptides | -0.011760 |
| 194 | COG0433 | COG0433 Predicted ATPase | -0.011728 |
| 195 | 33A63 | Psort location Cytoplasmic, score | -0.011715 |
| 196 | 2ZDT8 | 0.011714 | |
| 197 | COG2259 | Doxx family | -0.011705 |
| 198 | 31G01 | TniQ | 0.011704 |
| 199 | 337E4 | YvrJ protein family | 0.011699 |
| 200 | COG0423 | glycine-tRNA ligase activity | -0.011673 |
| 201 | COG1582 | Flagellar protein (FlbD) | -0.011657 |
| 202 | 2Z7K0 | 0.011631 | |
| 203 | 334IY | Domain of unknown function (DUF1844) | 0.011630 |
| 204 | 3312Z | 0.011629 | |
| 205 | COG0853 | Catalyzes the pyruvoyl-dependent decarboxylation of aspartate to produce beta-alanine | -0.011611 |
| 206 | 32WU5 | Class III cytochrome C family | 0.011601 |
| 207 | COG1600 | Catalyzes the conversion of epoxyqueuosine (oQ) to queuosine (Q), which is a hypermodified base found in the wobble positions of tRNA(Asp), tRNA(Asn), tRNA(His) and tRNA(Tyr) | -0.011588 |
| 208 | COG0684 | Catalyzes the aldol cleavage of 4-hydroxy-4-methyl-2- oxoglutarate (HMG) into 2 molecules of pyruvate. Also contains a secondary oxaloacetate (OAA) decarboxylase activity due to the common pyruvate enolate transition state formed following C-C bond cleavage in the retro-aldol and decarboxylation reactions | -0.011583 |
| 209 | COG1477 | protein flavinylation | -0.011573 |
| 210 | COG4594 | ABC-type Fe3 -citrate transport system, periplasmic component | -0.011560 |
| 211 | COG1606 | tRNA processing | 0.011549 |
| 212 | COG0046 | Part of the phosphoribosylformylglycinamidine synthase complex involved in the purines biosynthetic pathway. Catalyzes the ATP-dependent conversion of formylglycinamide ribonucleotide (FGAR) and glutamine to yield formylglycinamidine ribonucleotide (FGAM) and glutamate. The FGAM synthase complex is composed of three subunits. PurQ produces an ammonia molecule by converting glutamine to glutamate. PurL transfers the ammonia molecule to FGAR to form FGAM in an ATP-dependent manner. PurS interacts with PurQ and PurL and is thought to assist in the transfer of the ammonia molecule from PurQ to PurL | -0.011527 |
| 213 | COG1034 | ATP synthesis coupled electron transport | -0.011520 |
| 214 | 2Z852 | 0.011519 | |
| 215 | 31X2N | 0.011519 | |
| 216 | COG4303 | Ethanolamine ammonia lyase large subunit | 0.011505 |
| 217 | COG3316 | Transposase and inactivated derivatives | 0.011504 |
| 218 | COG0520 | Catalyzes the removal of elemental sulfur and selenium atoms from L-cysteine, L-cystine, L-selenocysteine, and L- selenocystine to produce L-alanine | -0.011475 |
| 219 | COG3944 | biosynthesis protein | 0.011464 |
| 220 | COG0422 | Catalyzes the synthesis of the hydroxymethylpyrimidine phosphate (HMP-P) moiety of thiamine from aminoimidazole ribotide (AIR) in a radical S-adenosyl-L-methionine (SAM)-dependent reaction | -0.011407 |
| 221 | 32SJ1 | 0.011393 | |
| 222 | COG1237 | beta-lactamase domain protein | 0.011393 |
| 223 | COG2818 | Glycosylase | 0.011388 |
| 224 | COG0334 | Belongs to the Glu Leu Phe Val dehydrogenases family | -0.011332 |
| 225 | 33AYK | -0.011323 | |
| 226 | COG4589 | Belongs to the CDS family | -0.011313 |
| 227 | 31H6C | Type I restriction enzyme R protein N terminus (HSDR_N) | 0.011310 |
| 228 | COG0328 | RNA-DNA hybrid ribonuclease activity | -0.011310 |
| 229 | COG2166 | iron-sulfur cluster assembly | -0.011309 |
| 230 | COG2358 | TRAP transporter, solute receptor (TAXI family | 0.011307 |
| 231 | COG3270 | Specifically methylates the cytosine at position 1407 (m5C1407) of 16S rRNA | -0.011306 |
| 232 | COG0624 | Catalyzes the hydrolysis of N-succinyl-L,L- diaminopimelic acid (SDAP), forming succinate and LL-2,6- diaminoheptanedioate (DAP), an intermediate involved in the bacterial biosynthesis of lysine and meso-diaminopimelic acid, an essential component of bacterial cell walls | -0.011291 |
| 233 | COG5416 | Lipopolysaccharide assembly protein A domain | 0.011284 |
| 234 | 33IJI | 0.011273 | |
| 235 | 3056W | 0.011266 | |
| 236 | COG2414 | aldehyde ferredoxin oxidoreductase | 0.011251 |
| 237 | 2ZAQ1 | Arylsulfotransferase Ig-like domain | 0.011250 |
| 238 | 33BFJ | 0.011239 | |
| 239 | 2ZQKN | 0.011231 | |
| 240 | COG3366 | ferrous iron transmembrane transporter activity | 0.011206 |
| 241 | COG2816 | NAD+ diphosphatase activity | -0.011193 |
| 242 | COG3862 | protein with conserved CXXC pairs | -0.011158 |
| 243 | 33ATC | Domain of unknown function (DUF4352) | 0.011156 |
| 244 | COG0407 | Catalyzes the decarboxylation of four acetate groups of uroporphyrinogen-III to yield coproporphyrinogen-III | -0.011150 |
| 245 | COG4147 | Belongs to the sodium solute symporter (SSF) (TC 2.A.21) family | 0.011130 |
| 246 | COG2369 | cell adhesion | 0.011120 |
| 247 | COG4676 | Protein conserved in bacteria | 0.011103 |
| 248 | COG3067 | Na( ) H( ) antiporter that extrudes sodium in exchange for external protons | 0.011093 |
| 249 | COG1234 | tRNA 3'-trailer cleavage | 0.011063 |
| 250 | 33T1G | 0.011057 |