Model Internals
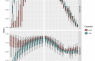
Each PhenDB model is trained on sets of bacterial ENOGs (orthologous groups from EggNOG 4.5), which have or have not been identified in the training genomes. Each ENOG is given a weight, with the magnitude of the weight being the importance of that ENOG for the final prediction. The sign of the weight indicates whether the presence (positive weight) or absence (negative weight) of this ENOG is indicative of the trait.
This table lists the 250 highest-ranking ENOGs of this model.
| rank in model | enog name | enog description | weight in model |
|---|---|---|---|
| 1 | COG1620 | l-lactate permease | 0.054525 |
| 2 | COG2837 | iron assimilation | 0.041748 |
| 3 | COG1199 | ATP-dependent helicase activity | -0.039120 |
| 4 | COG1283 | sodium-dependent phosphate transmembrane transporter activity | -0.028097 |
| 5 | COG3663 | DNA glycosylase | 0.026918 |
| 6 | COG4976 | Methyltransferase | -0.026884 |
| 7 | COG0315 | cyclic pyranopterin monophosphate synthase activity | 0.026807 |
| 8 | COG4753 | response regulator | 0.026794 |
| 9 | COG3239 | Fatty acid desaturase | -0.026624 |
| 10 | COG2314 | TM2 domain | -0.026469 |
| 11 | COG0371 | Dehydrogenase | -0.026319 |
| 12 | COG0510 | thiamine kinase activity | -0.026129 |
| 13 | COG2704 | Responsible for the transport of C4-dicarboxylates from the periplasm across the inner membrane | 0.026073 |
| 14 | COG2180 | nitrate reductase molybdenum cofactor assembly chaperone | 0.025707 |
| 15 | COG2040 | homocysteine | 0.025670 |
| 16 | COG1191 | sigma factor activity | -0.025617 |
| 17 | COG3620 | sequence-specific DNA binding | 0.025616 |
| 18 | COG4283 | Protein of unknown function (DUF1706) | -0.025599 |
| 19 | COG1275 | C4-dicarboxylate transporter malic acid transport protein | -0.025481 |
| 20 | COG0785 | Cytochrome C biogenesis protein | -0.025021 |
| 21 | COG2896 | Catalyzes the cyclization of GTP to (8S)-3',8-cyclo-7,8- dihydroguanosine 5'-triphosphate | 0.024985 |
| 22 | COG2015 | COG2015, Alkyl sulfatase and related hydrolases | -0.024911 |
| 23 | COG1887 | Glycosyl glycerophosphate transferases involved in teichoic acid biosynthesis TagF TagB EpsJ RodC | 0.024664 |
| 24 | COG0696 | Catalyzes the interconversion of 2-phosphoglycerate and 3-phosphoglycerate | 0.024636 |
| 25 | COG1600 | Catalyzes the conversion of epoxyqueuosine (oQ) to queuosine (Q), which is a hypermodified base found in the wobble positions of tRNA(Asp), tRNA(Asn), tRNA(His) and tRNA(Tyr) | -0.024351 |
| 26 | COG3591 | Belongs to the peptidase S1B family | -0.024337 |
| 27 | 3381Y | Antitoxin component of a toxin-antitoxin (TA) module | 0.024089 |
| 28 | COG2183 | response to ionizing radiation | -0.023836 |
| 29 | 2ZWWV | ORF located using Blastx | 0.023753 |
| 30 | COG1182 | Catalyzes the reductive cleavage of azo bond in aromatic azo compounds to the corresponding amines. Requires NADH, but not NADPH, as an electron donor for its activity | -0.023581 |
| 31 | COG0715 | COG0715 ABC-type nitrate sulfonate bicarbonate transport systems periplasmic components | -0.023532 |
| 32 | COG1951 | Catalyzes the reversible hydration of fumarate to (S)- malate | 0.023120 |
| 33 | COG3378 | Phage plasmid primase P4 family | 0.023075 |
| 34 | COG3376 | Belongs to the NiCoT transporter (TC 2.A.52) family | 0.023056 |
| 35 | COG3668 | Plasmid stabilization system | 0.022930 |
| 36 | COG0157 | Belongs to the NadC ModD family | -0.022717 |
| 37 | COG0741 | lytic transglycosylase activity | -0.022291 |
| 38 | COG1575 | Belongs to the MenA family. Type 1 subfamily | -0.022256 |
| 39 | COG1742 | UPF0060 membrane protein | 0.022229 |
| 40 | 3448G | 'Pfam Bacterial regulatory proteins, tetR family | -0.022222 |
| 41 | COG2879 | small protein | 0.022152 |
| 42 | COG1853 | conserved protein domain typically associated with flavoprotein oxygenases, DIM6 NTAB family | -0.022149 |
| 43 | COG1903 | Catalyzes the methylation of C-1 in cobalt-precorrin-5B to form cobalt-precorrin-6A | 0.021960 |
| 44 | COG0520 | Catalyzes the removal of elemental sulfur and selenium atoms from L-cysteine, L-cystine, L-selenocysteine, and L- selenocystine to produce L-alanine | -0.021869 |
| 45 | COG0400 | carboxylic ester hydrolase activity | -0.021824 |
| 46 | COG1546 | Belongs to the CinA family | -0.021447 |
| 47 | COG3475 | LICD family | 0.021197 |
| 48 | COG1129 | ABC transporter | -0.021192 |
| 49 | COG1010 | Tetrapyrrole (Corrin/Porphyrin) Methylases | 0.021128 |
| 50 | COG2336 | PFAM SpoVT AbrB | -0.020993 |
| 51 | COG2984 | ABC transporter substrate binding protein | 0.020968 |
| 52 | COG0753 | catalase activity | 0.020942 |
| 53 | COG4145 | pantothenate transmembrane transporter activity | -0.020893 |
| 54 | COG2073 | Cobalamin biosynthesis protein cbiG | 0.020867 |
| 55 | COG1211 | 2-C-methyl-D-erythritol 4-phosphate cytidylyltransferase activity | -0.020632 |
| 56 | COG0646 | methionine synthase | -0.020592 |
| 57 | COG3610 | response to peptide | 0.020544 |
| 58 | COG2056 | Na+-H+ antiporter family | 0.020436 |
| 59 | COG1140 | nitrate reductase beta subunit | 0.020398 |
| 60 | COG1364 | Catalyzes two activities which are involved in the cyclic version of arginine biosynthesis the synthesis of N- acetylglutamate from glutamate and acetyl-CoA as the acetyl donor, and of ornithine by transacetylation between N(2)-acetylornithine and glutamate | -0.020341 |
| 61 | COG3942 | CHAP domain | -0.020280 |
| 62 | COG4607 | ABC-type enterochelin transport system periplasmic component | -0.020267 |
| 63 | COG1301 | dicarboxylic acid transport | 0.020258 |
| 64 | COG0805 | protein transport | 0.020222 |
| 65 | COG4260 | virion core protein, lumpy skin disease virus | -0.020134 |
| 66 | COG0384 | phenazine biosynthesis protein PhzF | -0.020012 |
| 67 | COG2820 | Catalyzes the reversible phosphorylytic cleavage of uridine and deoxyuridine to uracil and ribose- or deoxyribose-1- phosphate. The produced molecules are then utilized as carbon and energy sources or in the rescue of pyrimidine bases for nucleotide synthesis | -0.019937 |
| 68 | COG3250 | beta-galactosidase activity | 0.019889 |
| 69 | COG3081 | Nucleoid-associated protein | 0.019760 |
| 70 | 2Z8PG | -0.019733 | |
| 71 | COG1321 | iron dependent repressor | -0.019617 |
| 72 | 33JI9 | PFAM FeoA family protein | -0.019580 |
| 73 | COG0717 | Belongs to the dCTP deaminase family | -0.019572 |
| 74 | COG1363 | Peptidase, m42 | -0.019565 |
| 75 | COG1826 | Part of the twin-arginine translocation (Tat) system that transports large folded proteins containing a characteristic twin-arginine motif in their signal peptide across membranes | 0.019555 |
| 76 | COG3174 | membrane | 0.019503 |
| 77 | COG3303 | Catalyzes the reduction of nitrite to ammonia, consuming six electrons in the process | 0.019341 |
| 78 | COG3426 | butyrate kinase activity | -0.019325 |
| 79 | COG4907 | membrane protein (DUF2207) | 0.019267 |
| 80 | COG2848 | UPF0210 protein | -0.019144 |
| 81 | COG4213 | ABC transporter substrate-binding protein | 0.019120 |
| 82 | COG5015 | Pyridoxamine 5'-phosphate oxidase | -0.019091 |
| 83 | COG3944 | biosynthesis protein | 0.019064 |
| 84 | COG2271 | Major facilitator Superfamily | 0.019007 |
| 85 | COG0076 | glutamate decarboxylase activity | -0.018999 |
| 86 | COG1006 | Multisubunit Na H antiporter MnhC subunit | -0.018968 |
| 87 | 33AEP | Domain of unknown function (DUF4430) | 0.018966 |
| 88 | COG0464 | growth | -0.018913 |
| 89 | COG4224 | UPF0291 protein | 0.018863 |
| 90 | COG4565 | transcriptional regulatory protein | -0.018804 |
| 91 | COG0523 | cobalamin synthesis protein | 0.018778 |
| 92 | COG4768 | protein containing a divergent version of the methyl-accepting chemotaxis-like domain | -0.018743 |
| 93 | COG2918 | glutathione biosynthetic process | -0.018726 |
| 94 | COG2181 | nitrate reductase activity | 0.018715 |
| 95 | COG0581 | inorganic phosphate transmembrane transporter activity | 0.018569 |
| 96 | COG1775 | 2-hydroxyglutaryl-CoA dehydratase, D-component | -0.018544 |
| 97 | COG1605 | Chorismate mutase | -0.018533 |
| 98 | COG3336 | cytochrome c oxidase | -0.018457 |
| 99 | COG1014 | Oxidoreductase required for the transfer of electrons from pyruvate to flavodoxin | -0.018453 |
| 100 | COG0398 | Pfam SNARE associated Golgi protein | -0.018449 |
| 101 | COG1143 | NDH-1 shuttles electrons from NADH, via FMN and iron- sulfur (Fe-S) centers, to quinones in the respiratory chain. The immediate electron acceptor for the enzyme in this species is believed to be ubiquinone. Couples the redox reaction to proton translocation (for every two electrons transferred, four hydrogen ions are translocated across the cytoplasmic membrane), and thus conserves the redox energy in a proton gradient | 0.018422 |
| 102 | COG1533 | DNA photolyase activity | 0.018384 |
| 103 | COG0616 | signal peptide processing | -0.018384 |
| 104 | COG0562 | UDP-galactopyranose mutase | -0.018370 |
| 105 | COG0657 | acetylesterase activity | 0.018346 |
| 106 | COG0610 | Subunit R is required for both nuclease and ATPase activities, but not for modification | 0.018315 |
| 107 | 30BKD | Bacterial regulatory proteins, lacI family | 0.018304 |
| 108 | COG4336 | lyase activity | -0.018245 |
| 109 | COG0145 | ligase activity | -0.018226 |
| 110 | COG1528 | Iron-storage protein | -0.018174 |
| 111 | COG3870 | Protein conserved in bacteria | 0.018158 |
| 112 | COG1178 | ABC-type Fe3 transport system permease component | -0.018155 |
| 113 | COG2184 | nucleotidyltransferase activity | -0.018130 |
| 114 | COG1085 | galactose-1-phosphate uridylyltransferase | 0.018093 |
| 115 | COG0427 | acetyl-CoA hydrolase | 0.018091 |
| 116 | COG2113 | Glycine betaine | 0.018031 |
| 117 | COG2199 | diguanylate cyclase activity | 0.018024 |
| 118 | COG2173 | Catalyzes hydrolysis of the D-alanyl-D-alanine dipeptide | 0.018007 |
| 119 | COG2875 | Belongs to the precorrin methyltransferase family | 0.017966 |
| 120 | 32Y15 | Psort location CytoplasmicMembrane, score | 0.017957 |
| 121 | COG3921 | Protein conserved in bacteria | 0.017941 |
| 122 | COG1135 | Part of the ABC transporter complex MetNIQ involved in methionine import. Responsible for energy coupling to the transport system | 0.017913 |
| 123 | COG3585 | molybdate ion transport | -0.017909 |
| 124 | 32STP | 0.017869 | |
| 125 | COG4540 | Baseplate assembly protein | -0.017856 |
| 126 | COG0859 | PFAM glycosyl transferase family 9 | -0.017854 |
| 127 | COG0346 | lactoylglutathione lyase activity | -0.017833 |
| 128 | COG5036 | VTC domain | -0.017804 |
| 129 | 308P1 | 0.017766 | |
| 130 | COG4720 | Psort location CytoplasmicMembrane, score | 0.017742 |
| 131 | COG1794 | racemase activity, acting on amino acids and derivatives | 0.017683 |
| 132 | COG1735 | metal-dependent hydrolase with the TIM-barrel fold | -0.017623 |
| 133 | COG3037 | sugar-specific permease SgaT UlaA | -0.017612 |
| 134 | COG3755 | Protein conserved in bacteria | 0.017487 |
| 135 | 33CG9 | GDSL-like Lipase/Acylhydrolase | 0.017456 |
| 136 | 32RPZ | Domain of unknown function (DUF4811) | 0.017450 |
| 137 | COG1443 | Catalyzes the 1,3-allylic rearrangement of the homoallylic substrate isopentenyl (IPP) to its highly electrophilic allylic isomer, dimethylallyl diphosphate (DMAPP) | -0.017428 |
| 138 | COG0553 | Transcription regulator that activates transcription by stimulating RNA polymerase (RNAP) recycling in case of stress conditions such as supercoiled DNA or high salt concentrations. Probably acts by releasing the RNAP, when it is trapped or immobilized on tightly supercoiled DNA. Does not activate transcription on linear DNA. Probably not involved in DNA repair | -0.017397 |
| 139 | COG2084 | Dehydrogenase | -0.017385 |
| 140 | COG3440 | Restriction endonuclease | -0.017362 |
| 141 | COG2365 | Tyrosine phosphatase family | 0.017295 |
| 142 | COG3945 | hemerythrin HHE cation binding domain | -0.017288 |
| 143 | COG4289 | Protein conserved in bacteria | 0.017281 |
| 144 | 32T36 | 0.017231 | |
| 145 | 33CM8 | 0.017184 | |
| 146 | COG4454 | pyrroloquinoline quinone binding | 0.017173 |
| 147 | COG2972 | Histidine kinase | -0.017093 |
| 148 | COG0295 | cytidine deaminase activity | -0.017083 |
| 149 | COG0379 | Catalyzes the condensation of iminoaspartate with dihydroxyacetone phosphate to form quinolinate | -0.017072 |
| 150 | COG1954 | Regulates expression of the glpD operon. In the presence of glycerol 3-phosphate (G3P) causes antitermination of transcription of glpD at the inverted repeat of the leader region to enhance its transcription. Binds and stabilizes glpD leader mRNA | 0.017065 |
| 151 | 31W2T | Universal stress protein family | -0.017065 |
| 152 | COG1804 | L-carnitine dehydratase bile acid-inducible protein F | -0.017064 |
| 153 | COG1937 | Protein conserved in bacteria | 0.017047 |
| 154 | COG2329 | Antibiotic biosynthesis monooxygenase | -0.017036 |
| 155 | COG0521 | Mo-molybdopterin cofactor biosynthetic process | 0.017035 |
| 156 | COG2368 | 4-hydroxyphenylacetate | -0.017034 |
| 157 | COG3682 | Transcriptional regulator | -0.017030 |
| 158 | COG0780 | Catalyzes the NADPH-dependent reduction of 7-cyano-7- deazaguanine (preQ0) to 7-aminomethyl-7-deazaguanine (preQ1) | 0.017015 |
| 159 | COG2227 | 3-demethylubiquinone-9 3-O-methyltransferase activity | 0.017007 |
| 160 | COG0341 | Part of the Sec protein translocase complex. Interacts with the SecYEG preprotein conducting channel. SecDF uses the proton motive force (PMF) to complete protein translocation after the ATP-dependent function of SecA | -0.016990 |
| 161 | COG2011 | ABC-type metal ion transport system permease component | 0.016968 |
| 162 | COG0112 | Catalyzes the reversible interconversion of serine and glycine with tetrahydrofolate (THF) serving as the one-carbon carrier. This reaction serves as the major source of one-carbon groups required for the biosynthesis of purines, thymidylate, methionine, and other important biomolecules. Also exhibits THF- independent aldolase activity toward beta-hydroxyamino acids, producing glycine and aldehydes, via a retro-aldol mechanism | 0.016935 |
| 163 | 32A7C | GDSL-like Lipase/Acylhydrolase family | 0.016912 |
| 164 | COG2826 | transposase and inactivated derivatives, IS30 family | 0.016900 |
| 165 | COG0560 | Phosphoserine phosphatase | -0.016819 |
| 166 | COG1012 | belongs to the aldehyde dehydrogenase family | -0.016819 |
| 167 | COG3973 | AAA domain | -0.016803 |
| 168 | COG4670 | ketone body catabolic process | -0.016798 |
| 169 | COG4185 | zeta toxin | -0.016772 |
| 170 | COG3093 | addiction module antidote protein HigA | 0.016707 |
| 171 | COG4464 | protein tyrosine phosphatase activity | 0.016695 |
| 172 | COG0590 | tRNA wobble adenosine to inosine editing | -0.016677 |
| 173 | COG1410 | methionine synthase | 0.016673 |
| 174 | COG2030 | dehydratase | -0.016656 |
| 175 | COG0605 | Destroys radicals which are normally produced within the cells and which are toxic to biological systems | -0.016618 |
| 176 | COG1764 | response to oxidative stress | 0.016599 |
| 177 | COG1186 | Peptide chain release factor 2 directs the termination of translation in response to the peptide chain termination codons UGA and UAA | 0.016575 |
| 178 | COG1104 | cysteine desulfurase activity | -0.016572 |
| 179 | COG2898 | Catalyzes the transfer of a lysyl group from L-lysyl- tRNA(Lys) to membrane-bound phosphatidylglycerol (PG), which produces lysylphosphatidylglycerol (LPG), a major component of the bacterial membrane with a positive net charge. LPG synthesis contributes to bacterial virulence as it is involved in the resistance mechanism against cationic antimicrobial peptides (CAMP) produces by the host's immune system (defensins, cathelicidins) and by the competing microorganisms | 0.016494 |
| 180 | 32G2Z | Lysin motif | 0.016493 |
| 181 | COG5578 | integral membrane protein | -0.016441 |
| 182 | COG1442 | glycosyl transferase family 8 | 0.016428 |
| 183 | COG3059 | membrane | 0.016416 |
| 184 | 33J2G | Control of competence regulator ComK, YlbF/YmcA | 0.016407 |
| 185 | 2Z948 | Psort location CytoplasmicMembrane, score 10.00 | -0.016362 |
| 186 | COG3039 | Transposase | 0.016346 |
| 187 | COG1477 | protein flavinylation | -0.016264 |
| 188 | COG0608 | single-stranded DNA 5'-3' exodeoxyribonuclease activity | -0.016258 |
| 189 | COG1989 | Cleaves type-4 fimbrial leader sequence and methylates the N-terminal (generally Phe) residue | 0.016246 |
| 190 | COG3851 | Histidine kinase | 0.016156 |
| 191 | COG4109 | DRTGG domain | -0.016150 |
| 192 | COG4555 | ABC transporter | -0.016120 |
| 193 | COG2706 | 6-phosphogluconolactonase activity | -0.016095 |
| 194 | COG0027 | Involved in the de novo purine biosynthesis. Catalyzes the transfer of formate to 5-phospho-ribosyl-glycinamide (GAR), producing 5-phospho-ribosyl-N-formylglycinamide (FGAR). Formate is provided by PurU via hydrolysis of 10-formyl-tetrahydrofolate | -0.016084 |
| 195 | COG3822 | ABC-type sugar transport system, auxiliary component | -0.016081 |
| 196 | COG2856 | Zn peptidase | -0.016080 |
| 197 | COG4403 | Lanthionine synthetase C family protein | -0.016069 |
| 198 | COG5614 | head-tail adaptor | 0.016050 |
| 199 | COG3328 | transposase activity | 0.016036 |
| 200 | 32QT1 | -0.016015 | |
| 201 | COG0106 | 1-(5-phosphoribosyl)-5- 5-phosphoribosylamino)methylideneamino imidazole-4-carboxamide isomerase | -0.015973 |
| 202 | COG0704 | Plays a role in the regulation of phosphate uptake | 0.015972 |
| 203 | COG2035 | Membrane | -0.015951 |
| 204 | 32W8Q | 0.015949 | |
| 205 | COG4641 | Protein conserved in bacteria | 0.015933 |
| 206 | 33D6A | Nucleotidyltransferase substrate binding protein like | 0.015927 |
| 207 | COG1070 | Carbohydrate kinase | 0.015895 |
| 208 | COG0814 | amino acid | 0.015888 |
| 209 | COG2177 | Part of the ABC transporter FtsEX involved in | -0.015823 |
| 210 | COG0122 | 3-methyladenine DNA glycosylase 8-oxoguanine DNA glycosylase | -0.015816 |
| 211 | COG1715 | Restriction endonuclease | -0.015783 |
| 212 | COG4615 | Cyclic peptide transporter | 0.015783 |
| 213 | COG0314 | molybdopterin synthase activity | 0.015758 |
| 214 | COG2976 | Protein conserved in bacteria | -0.015755 |
| 215 | COG3369 | Iron-binding zinc finger CDGSH type | -0.015733 |
| 216 | COG1007 | ATP synthesis coupled electron transport | 0.015643 |
| 217 | COG3981 | Acetyltransferase (GNAT) domain | -0.015620 |
| 218 | COG0671 | phosphoesterase, PA-phosphatase related | -0.015619 |
| 219 | COG2352 | Forms oxaloacetate, a four-carbon dicarboxylic acid source for the tricarboxylic acid cycle | -0.015614 |
| 220 | COG0662 | Cupin 2, conserved barrel domain protein | 0.015613 |
| 221 | COG1171 | Catalyzes the anaerobic formation of alpha-ketobutyrate and ammonia from threonine in a two-step reaction. The first step involved a dehydration of threonine and a production of enamine intermediates (aminocrotonate), which tautomerizes to its imine form (iminobutyrate). Both intermediates are unstable and short- lived. The second step is the nonenzymatic hydrolysis of the enamine imine intermediates to form 2-ketobutyrate and free ammonia. In the low water environment of the cell, the second step is accelerated by RidA | -0.015610 |
| 222 | COG1464 | Belongs to the NlpA lipoprotein family | 0.015595 |
| 223 | COG1556 | Lactate utilization protein | 0.015541 |
| 224 | COG2963 | transposase activity | 0.015525 |
| 225 | COG0820 | rRNA (adenine-C2-)-methyltransferase activity | -0.015517 |
| 226 | COG1484 | DNA-dependent DNA replication | 0.015501 |
| 227 | COG2370 | cobalamin-transporting ATPase activity | 0.015477 |
| 228 | 2ZGY7 | 0.015456 | |
| 229 | COG4465 | DNA-binding protein that represses the expression of many genes that are induced as cells make the transition from rapid exponential growth to stationary phase. It is a GTP-binding protein that senses the intracellular GTP concentration as an indicator of nutritional limitations. At low GTP concentration it no longer binds GTP and stop to act as a transcriptional repressor | -0.015439 |
| 230 | COG4485 | Bacterial membrane protein, YfhO | 0.015432 |
| 231 | COG0645 | Protein conserved in bacteria | -0.015410 |
| 232 | COG1177 | transmembrane transport | 0.015406 |
| 233 | COG0474 | ATPase, P-type transporting, HAD superfamily, subfamily IC | -0.015386 |
| 234 | 3058S | 0.015355 | |
| 235 | COG4822 | anaerobic cobalamin biosynthetic process | 0.015338 |
| 236 | COG1062 | S-(hydroxymethyl)glutathione dehydrogenase activity | -0.015337 |
| 237 | 30C2I | -0.015335 | |
| 238 | COG2143 | COG2143 Thioredoxin-related protein | -0.015331 |
| 239 | COG0175 | sulfate reduction | 0.015331 |
| 240 | 346DU | GDSL-like Lipase/Acylhydrolase | 0.015325 |
| 241 | COG1785 | Belongs to the alkaline phosphatase family | -0.015325 |
| 242 | 33YTS | -0.015274 | |
| 243 | COG5457 | Domain of unknown function (DUF1127) | 0.015271 |
| 244 | COG0385 | Bile acid | 0.015268 |
| 245 | COG2884 | Cell division ATP-binding protein ftsE | -0.015265 |
| 246 | 30584 | 0.015260 | |
| 247 | COG3883 | PFAM NLP P60 protein | -0.015242 |
| 248 | COG4129 | membrane | -0.015239 |
| 249 | COG1979 | alcohol dehydrogenase | -0.015194 |
| 250 | COG2515 | 1-aminocyclopropane-1-carboxylate deaminase | 0.015190 |