Model Internals
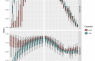
Each PhenDB model is trained on sets of bacterial ENOGs (orthologous groups from EggNOG 4.5), which have or have not been identified in the training genomes. Each ENOG is given a weight, with the magnitude of the weight being the importance of that ENOG for the final prediction. The sign of the weight indicates whether the presence (positive weight) or absence (negative weight) of this ENOG is indicative of the trait.
This table lists the 250 highest-ranking ENOGs of this model.
| rank in model | enog name | enog description | weight in model |
|---|---|---|---|
| 1 | COG1030 | water channel activity | 0.023160 |
| 2 | COG1613 | Sulfate ABC transporter periplasmic sulfate-binding protein | -0.023084 |
| 3 | COG4967 | type IV pilus modification protein PilV | 0.022022 |
| 4 | COG3505 | Type IV secretory pathway VirD4 | -0.019951 |
| 5 | COG1982 | Orn Lys Arg decarboxylase major | -0.019714 |
| 6 | COG0644 | oxidoreductase | 0.019600 |
| 7 | COG0687 | Required for the activity of the bacterial periplasmic transport system of putrescine | -0.019478 |
| 8 | COG4798 | methyltransferase | 0.019001 |
| 9 | COG1610 | YqeY-like protein | 0.018992 |
| 10 | COG1027 | Aspartate ammonia-lyase | -0.018969 |
| 11 | COG0738 | Major facilitator superfamily | -0.018681 |
| 12 | COG1292 | Belongs to the BCCT transporter (TC 2.A.15) family | 0.018640 |
| 13 | COG4208 | sulfate ABC transporter | -0.018362 |
| 14 | COG0698 | ribose 5-phosphate isomerase | -0.017987 |
| 15 | COG4446 | Protein conserved in bacteria | 0.017954 |
| 16 | COG3806 | Anti-sigma factor | 0.017829 |
| 17 | COG0655 | NAD(P)H dehydrogenase (quinone) activity | 0.017825 |
| 18 | COG0555 | PFAM binding-protein-dependent transport systems inner membrane component | -0.017576 |
| 19 | COG1912 | Pfam S-adenosyl-l-methionine hydroxide adenosyltransferase | 0.017543 |
| 20 | COG0340 | biotin-[acetyl-CoA-carboxylase] ligase activity | -0.017412 |
| 21 | COG1515 | deoxyribonuclease V activity | 0.017075 |
| 22 | COG3090 | Trap-type c4-dicarboxylate transport system, small permease component | 0.016968 |
| 23 | COG1528 | Iron-storage protein | -0.016890 |
| 24 | COG1089 | Catalyzes the conversion of GDP-D-mannose to GDP-4- dehydro-6-deoxy-D-mannose | -0.016859 |
| 25 | COG1983 | PspC domain protein | 0.016832 |
| 26 | COG1525 | nuclease | -0.016797 |
| 27 | COG1482 | cell wall glycoprotein biosynthetic process | -0.016757 |
| 28 | COG1233 | COG1233 Phytoene dehydrogenase and related | 0.016730 |
| 29 | COG0469 | pyruvate kinase activity | -0.016659 |
| 30 | COG4559 | Part of the ABC transporter complex HmuTUV involved in hemin import. Responsible for energy coupling to the transport system | 0.016632 |
| 31 | 2Z7ZD | Ion transport protein | 0.016522 |
| 32 | COG1272 | protein, Hemolysin III | 0.016488 |
| 33 | COG0123 | including yeast histone deacetylase and acetoin utilization protein | 0.016383 |
| 34 | COG2005 | Transcriptional regulator | -0.016250 |
| 35 | COG0390 | transport system, permease component | 0.016240 |
| 36 | COG3769 | mannosylglycerate metabolic process | 0.015683 |
| 37 | COG2060 | Part of the high-affinity ATP-driven potassium transport (or Kdp) system, which catalyzes the hydrolysis of ATP coupled with the electrogenic transport of potassium into the cytoplasm. This subunit binds and transports the potassium across the cytoplasmic membrane | -0.015676 |
| 38 | COG2156 | Part of the high-affinity ATP-driven potassium transport (or Kdp) system, which catalyzes the hydrolysis of ATP coupled with the electrogenic transport of potassium into the cytoplasm. This subunit acts as a catalytic chaperone that increases the ATP- binding affinity of the ATP-hydrolyzing subunit KdpB by the formation of a transient KdpB KdpC ATP ternary complex | -0.015530 |
| 39 | COG4178 | (ABC) transporter | -0.015504 |
| 40 | 2ZBMT | Protein of unknown function (DUF2959) | 0.015492 |
| 41 | COG4325 | translation initiation factor activity | 0.015423 |
| 42 | COG3414 | Phosphotransferase system galactitol-specific IIB component | -0.015308 |
| 43 | COG2216 | Part of the high-affinity ATP-driven potassium transport (or Kdp) system, which catalyzes the hydrolysis of ATP coupled with the electrogenic transport of potassium into the cytoplasm. This subunit is responsible for energy coupling to the transport system | -0.015220 |
| 44 | COG4993 | Dehydrogenase | 0.015185 |
| 45 | COG3591 | Belongs to the peptidase S1B family | 0.015167 |
| 46 | COG2948 | multi-organism process | -0.015111 |
| 47 | COG0393 | Putative heavy-metal-binding | 0.015028 |
| 48 | COG2721 | sulfolactate sulfo-lyase activity | -0.014984 |
| 49 | COG0657 | acetylesterase activity | -0.014944 |
| 50 | COG4666 | Tripartite ATP-independent periplasmic transporter, DctM component | 0.014865 |
| 51 | COG0822 | iron-sulfur transferase activity | -0.014826 |
| 52 | COG5006 | permease, DMT superfamily | 0.014757 |
| 53 | COG0692 | Excises uracil residues from the DNA which can arise as a result of misincorporation of dUMP residues by DNA polymerase or due to deamination of cytosine | -0.014754 |
| 54 | COG4717 | serine-type aminopeptidase activity | 0.014751 |
| 55 | COG2945 | thiolester hydrolase activity | -0.014708 |
| 56 | COG0622 | Phosphoesterase | -0.014579 |
| 57 | COG4261 | Acyltransferase | -0.014555 |
| 58 | COG2271 | Major facilitator Superfamily | -0.014449 |
| 59 | COG4536 | flavin adenine dinucleotide binding | 0.014394 |
| 60 | 33ARA | COG0526, thiol-disulfide isomerase and thioredoxins | 0.014391 |
| 61 | COG1914 | H( )-stimulated, divalent metal cation uptake system | -0.014341 |
| 62 | COG4928 | KAP family P-loop domain | 0.014340 |
| 63 | COG2128 | Antioxidant protein with alkyl hydroperoxidase activity. Required for the reduction of the AhpC active site cysteine residues and for the regeneration of the AhpC enzyme activity | 0.014321 |
| 64 | 32RYR | Domain of unknown function (DUF4112) | 0.014290 |
| 65 | COG0530 | calcium, potassium:sodium antiporter activity | 0.014250 |
| 66 | COG1654 | Acts both as a biotin-- acetyl-CoA-carboxylase ligase and a biotin-operon repressor. In the presence of ATP, BirA activates biotin to form the BirA-biotinyl-5'-adenylate (BirA-bio- 5'-AMP or holoBirA) complex. HoloBirA can either transfer the biotinyl moiety to the biotin carboxyl carrier protein (BCCP) subunit of acetyl-CoA carboxylase, or bind to the biotin operator site and inhibit transcription of the operon | 0.014230 |
| 67 | COG0796 | Provides the (R)-glutamate required for cell wall biosynthesis | -0.014207 |
| 68 | COG2733 | membrane | -0.014192 |
| 69 | COG1490 | D-aminoacyl-tRNA deacylase activity | 0.014167 |
| 70 | COG1063 | alcohol dehydrogenase | 0.014026 |
| 71 | COG3119 | arylsulfatase activity | -0.013954 |
| 72 | COG1162 | GTPase activity | -0.013777 |
| 73 | COG1502 | Catalyzes the reversible phosphatidyl group transfer from one phosphatidylglycerol molecule to another to form cardiolipin (CL) (diphosphatidylglycerol) and glycerol | -0.013714 |
| 74 | COG3451 | type IV secretory pathway VirB4 | -0.013660 |
| 75 | COG1732 | Glycine betaine | -0.013603 |
| 76 | COG1970 | mechanosensitive ion channel activity | -0.013512 |
| 77 | COG1035 | Coenzyme F420 hydrogenase/dehydrogenase, beta subunit C terminus | 0.013506 |
| 78 | COG1337 | RAMP superfamily | 0.013490 |
| 79 | COG4104 | PAAR repeat-containing protein | -0.013416 |
| 80 | COG1550 | Protein conserved in bacteria | 0.013401 |
| 81 | COG3707 | response regulator | -0.013320 |
| 82 | COG1462 | curli production assembly transport component CsgG | 0.013291 |
| 83 | COG1585 | Membrane protein implicated in regulation of membrane protease activity | -0.013257 |
| 84 | COG1074 | ATP-dependent DNA helicase activity | -0.013249 |
| 85 | COG0672 | )-iron permease | -0.013247 |
| 86 | COG1752 | Esterase of the alpha-beta hydrolase superfamily | 0.013237 |
| 87 | COG0466 | ATP-dependent serine protease that mediates the selective degradation of mutant and abnormal proteins as well as certain short-lived regulatory proteins. Required for cellular homeostasis and for survival from DNA damage and developmental changes induced by stress. Degrades polypeptides processively to yield small peptide fragments that are 5 to 10 amino acids long. Binds to DNA in a double-stranded, site-specific manner | -0.013194 |
| 88 | 32RM0 | 23S rRNA-intervening sequence protein | 0.013159 |
| 89 | COG0508 | dehydrogenase complex catalyzes the overall conversion of | -0.013096 |
| 90 | COG0626 | cystathionine gamma-synthase activity | -0.013048 |
| 91 | COG4912 | Dna alkylation repair | 0.012949 |
| 92 | COG1621 | beta-fructofuranosidase activity | -0.012945 |
| 93 | COG2859 | Protein conserved in bacteria | 0.012881 |
| 94 | COG1624 | Catalyzes the condensation of 2 ATP molecules into cyclic di-AMP (c-di-AMP), a second messenger used to regulate differing processes in different bacteria | -0.012835 |
| 95 | COG5305 | Membrane | -0.012816 |
| 96 | COG2234 | aminopeptidase activity | 0.012804 |
| 97 | COG0179 | Fumarylacetoacetate (FAA) hydrolase | 0.012796 |
| 98 | COG3193 | protein, possibly involved in utilization of glycolate and propanediol | -0.012791 |
| 99 | COG1324 | tolerance protein | 0.012751 |
| 100 | COG1284 | Uncharacterised 5xTM membrane BCR, YitT family COG1284 | -0.012681 |
| 101 | COG3415 | Transposase | 0.012647 |
| 102 | COG3453 | Protein conserved in bacteria | 0.012647 |
| 103 | COG0639 | Hydrolyzes diadenosine 5',5'''-P1,P4-tetraphosphate to yield ADP | 0.012643 |
| 104 | COG4103 | Tellurite resistance protein TerB | 0.012609 |
| 105 | COG5652 | VanZ like family | 0.012560 |
| 106 | COG1942 | 4-Oxalocrotonate Tautomerase | -0.012555 |
| 107 | COG0562 | UDP-galactopyranose mutase | -0.012548 |
| 108 | COG0520 | Catalyzes the removal of elemental sulfur and selenium atoms from L-cysteine, L-cystine, L-selenocysteine, and L- selenocystine to produce L-alanine | -0.012504 |
| 109 | COG1003 | The glycine cleavage system catalyzes the degradation of glycine. The P protein binds the alpha-amino group of glycine through its pyridoxal phosphate cofactor | 0.012451 |
| 110 | COG1757 | Na H antiporter | 0.012431 |
| 111 | COG1509 | lysine 2,3-aminomutase activity | 0.012418 |
| 112 | COG1975 | Xanthine and CO dehydrogenases maturation factor, XdhC CoxF family | -0.012403 |
| 113 | COG0241 | D,D-heptose 1,7-bisphosphate phosphatase | 0.012374 |
| 114 | COG3640 | PFAM CobQ CobB MinD ParA nucleotide binding domain | 0.012347 |
| 115 | COG1236 | Exonuclease of the beta-lactamase fold involved in RNA processing | 0.012347 |
| 116 | COG2826 | transposase and inactivated derivatives, IS30 family | -0.012334 |
| 117 | COG4132 | (ABC) transporter, permease | -0.012317 |
| 118 | COG1638 | TRAP-type C4-dicarboxylate transport system periplasmic component | 0.012280 |
| 119 | COG2501 | S4 domain | -0.012274 |
| 120 | COG0733 | neurotransmitter:sodium symporter activity | 0.012232 |
| 121 | COG1489 | DNA binding | 0.012212 |
| 122 | COG1607 | acyl-coa hydrolase | 0.012193 |
| 123 | COG4782 | Protein conserved in bacteria | 0.012190 |
| 124 | COG1141 | Ferredoxin | -0.012178 |
| 125 | COG4105 | Part of the outer membrane protein assembly complex, which is involved in assembly and insertion of beta-barrel proteins into the outer membrane | -0.012166 |
| 126 | COG0737 | Belongs to the 5'-nucleotidase family | -0.012157 |
| 127 | COG0432 | Pfam Uncharacterised protein family UPF0047 | 0.012124 |
| 128 | COG3158 | transport of potassium into the cell | -0.012107 |
| 129 | COG3245 | cytochrome | 0.012076 |
| 130 | COG0500 | methyltransferase | 0.012068 |
| 131 | COG1652 | LysM domain | -0.012064 |
| 132 | COG4487 | mechanosensitive ion channel activity | -0.012036 |
| 133 | COG2137 | regulation of response to DNA damage stimulus | 0.012033 |
| 134 | COG1763 | Mo-molybdopterin cofactor biosynthetic process | -0.012023 |
| 135 | COG1353 | crispr-associated protein | 0.012019 |
| 136 | COG4537 | Required for transformation and DNA binding | 0.011943 |
| 137 | COG2252 | PERMEase | -0.011927 |
| 138 | COG3504 | Conjugal transfer protein | -0.011860 |
| 139 | COG0027 | Involved in the de novo purine biosynthesis. Catalyzes the transfer of formate to 5-phospho-ribosyl-glycinamide (GAR), producing 5-phospho-ribosyl-N-formylglycinamide (FGAR). Formate is provided by PurU via hydrolysis of 10-formyl-tetrahydrofolate | -0.011855 |
| 140 | COG0204 | Acyl-transferase | -0.011849 |
| 141 | COG0566 | Belongs to the class IV-like SAM-binding methyltransferase superfamily. RNA methyltransferase TrmH family | -0.011849 |
| 142 | COG0656 | L-ascorbic acid biosynthetic process | -0.011798 |
| 143 | COG1893 | Catalyzes the NADPH-dependent reduction of ketopantoate into pantoic acid | -0.011779 |
| 144 | COG2968 | Membrane | 0.011775 |
| 145 | COG4327 | Domain of unknown function (DUF4212) | 0.011704 |
| 146 | COG3576 | pyridoxamine 5-phosphate | 0.011695 |
| 147 | COG3450 | Protein of unknown function (DUF861) | -0.011666 |
| 148 | COG2514 | catechol 2,3-dioxygenase activity | 0.011657 |
| 149 | COG2366 | antibiotic biosynthetic process | 0.011646 |
| 150 | COG3919 | ATP-grasp | 0.011627 |
| 151 | COG2738 | Putative neutral zinc metallopeptidase | 0.011622 |
| 152 | COG1918 | ferrous iron import across plasma membrane | -0.011541 |
| 153 | 32VXQ | NUDIX domain | 0.011541 |
| 154 | COG5639 | conserved small protein | -0.011536 |
| 155 | COG1122 | ATPase activity | -0.011526 |
| 156 | COG2190 | (PTS) system | -0.011510 |
| 157 | COG0609 | Belongs to the binding-protein-dependent transport system permease family. FecCD subfamily | 0.011510 |
| 158 | COG0791 | NLP P60 protein | -0.011473 |
| 159 | COG1522 | sequence-specific DNA binding | 0.011459 |
| 160 | COG1725 | Transcriptional regulator | -0.011448 |
| 161 | COG0699 | ATPase. Has a role at an early stage in the morphogenesis of the spore coat | 0.011366 |
| 162 | COG0707 | Cell wall formation. Catalyzes the transfer of a GlcNAc subunit on undecaprenyl-pyrophosphoryl-MurNAc-pentapeptide (lipid intermediate I) to form undecaprenyl-pyrophosphoryl-MurNAc- (pentapeptide)GlcNAc (lipid intermediate II) | -0.011362 |
| 163 | 32S6W | 0.011348 | |
| 164 | COG1432 | Conserved Protein | 0.011307 |
| 165 | COG3758 | protein conserved in bacteria | -0.011286 |
| 166 | COG3228 | Belongs to the MtfA family | 0.011285 |
| 167 | COG0435 | Glutathione S-transferase | 0.011247 |
| 168 | COG0306 | phosphate transporter | -0.011244 |
| 169 | COG3671 | membrane | 0.011236 |
| 170 | COG1011 | Hydrolase | 0.011227 |
| 171 | COG5551 | CRISPR-associated protein Cas6 | 0.011226 |
| 172 | COG1956 | GAF domain-containing protein | 0.011178 |
| 173 | COG2893 | phosphoenolpyruvate-dependent sugar phosphotransferase system | -0.011175 |
| 174 | COG1514 | Hydrolyzes RNA 2',3'-cyclic phosphodiester to an RNA 2'- phosphomonoester | 0.011153 |
| 175 | COG2081 | HI0933 family | 0.011145 |
| 176 | COG1720 | tRNA m6t6A37 methyltransferase activity | 0.011140 |
| 177 | COG3263 | regulation of cellular component size | 0.011138 |
| 178 | COG2975 | iron-sulfur cluster assembly | -0.011135 |
| 179 | COG4619 | ABC transporter | 0.011127 |
| 180 | COG2331 | Regulatory protein, FmdB family | 0.011126 |
| 181 | COG0255 | Belongs to the universal ribosomal protein uL29 family | -0.011106 |
| 182 | COG1780 | Probably involved in ribonucleotide reductase function | -0.011103 |
| 183 | COG3860 | Transcriptional regulator | 0.011097 |
| 184 | COG0314 | molybdopterin synthase activity | -0.011077 |
| 185 | COG0670 | Belongs to the BI1 family | -0.011076 |
| 186 | COG2509 | 5-formyltetrahydrofolate cyclo-ligase activity | -0.011069 |
| 187 | COG2358 | TRAP transporter, solute receptor (TAXI family | 0.011043 |
| 188 | COG2864 | formate dehydrogenase | -0.011035 |
| 189 | COG0218 | Necessary for normal cell division and for the maintenance of normal septation | -0.011014 |
| 190 | COG2361 | Protein of unknown function DUF86 | 0.010966 |
| 191 | COG2002 | toxin-antitoxin pair type II binding | 0.010950 |
| 192 | COG2907 | Flavin containing amine oxidoreductase | 0.010919 |
| 193 | COG3464 | Transposase | 0.010898 |
| 194 | COG0017 | Asparaginyl-tRNA synthetase | -0.010892 |
| 195 | 3279X | protein transport | 0.010889 |
| 196 | COG0549 | Belongs to the carbamate kinase family | -0.010888 |
| 197 | 31W2T | Universal stress protein family | 0.010876 |
| 198 | 32YVH | Bacterial inner membrane protein | 0.010872 |
| 199 | COG1505 | serine-type exopeptidase activity | 0.010847 |
| 200 | COG3956 | TIGRFAM MazG family protein | 0.010824 |
| 201 | 338CY | 0.010820 | |
| 202 | COG3378 | Phage plasmid primase P4 family | 0.010812 |
| 203 | COG3861 | electron transporter, transferring electrons within the cyclic electron transport pathway of photosynthesis activity | 0.010794 |
| 204 | COG1189 | Ribosomal RNA methyltransferase RrmJ FtsJ | -0.010777 |
| 205 | COG3149 | Involved in a type II secretion system (T2SS, formerly general secretion pathway, GSP) for the export of proteins | 0.010758 |
| 206 | COG0212 | Belongs to the 5-formyltetrahydrofolate cyclo-ligase family | -0.010753 |
| 207 | COG1497 | Winged helix-turn-helix DNA-binding | 0.010751 |
| 208 | COG4538 | SnoaL-like domain | 0.010729 |
| 209 | COG0560 | Phosphoserine phosphatase | -0.010701 |
| 210 | COG3568 | Endonuclease Exonuclease Phosphatase | 0.010698 |
| 211 | COG3809 | Transcription factor zinc-finger | 0.010688 |
| 212 | COG5377 | YqaJ viral recombinase family | -0.010661 |
| 213 | COG2254 | TIGRFAM CRISPR-associated helicase Cas3 | -0.010658 |
| 214 | COG4970 | Tfp pilus assembly protein FimT | 0.010653 |
| 215 | COG4395 | Tim44 | -0.010630 |
| 216 | COG5505 | Protein of unknown function (DUF819) | 0.010623 |
| 217 | COG1208 | COG1208 Nucleoside-diphosphate-sugar pyrophosphorylase involved in lipopolysaccharide biosynthesis translation initiation factor 2B, gamma epsilon subunits eIF-2Bgamma eIF-2Bepsilon | 0.010606 |
| 218 | COG1445 | protein-N(PI)-phosphohistidine-fructose phosphotransferase system transporter activity | -0.010597 |
| 219 | COG0611 | Catalyzes the ATP-dependent phosphorylation of thiamine- monophosphate (TMP) to form thiamine-pyrophosphate (TPP), the active form of vitamin B1 | 0.010578 |
| 220 | COG2519 | Catalyzes the S-adenosyl-L-methionine-dependent formation of N(1)-methyladenine at position 58 (m1A58) in tRNA | -0.010572 |
| 221 | COG4147 | Belongs to the sodium solute symporter (SSF) (TC 2.A.21) family | 0.010568 |
| 222 | COG0619 | Transmembrane (T) component of an energy-coupling factor (ECF) ABC-transporter complex. Unlike classic ABC transporters this ECF transporter provides the energy necessary to transport a number of different substrates | -0.010564 |
| 223 | COG0579 | oxidoreductase | -0.010536 |
| 224 | COG1127 | ABC transporter | -0.010521 |
| 225 | COG0003 | Pfam Anion-transporting ATPase | 0.010509 |
| 226 | COG0161 | adenosylmethionine-8-amino-7-oxononanoate transaminase activity | -0.010505 |
| 227 | COG5592 | hemerythrin HHE cation binding domain | -0.010492 |
| 228 | COG2017 | converts alpha-aldose to the beta-anomer | -0.010464 |
| 229 | COG3585 | molybdate ion transport | -0.010453 |
| 230 | COG3492 | Protein of unknown function (DUF1244) | 0.010443 |
| 231 | COG3137 | Salt-induced outer membrane protein | 0.010427 |
| 232 | COG5519 | DNA primase | 0.010410 |
| 233 | COG3069 | C4-dicarboxylate transmembrane transporter activity | -0.010401 |
| 234 | COG1115 | amino acid carrier protein | 0.010378 |
| 235 | COG0631 | protein serine/threonine phosphatase activity | -0.010324 |
| 236 | COG0282 | Catalyzes the formation of acetyl phosphate from acetate and ATP. Can also catalyze the reverse reaction | -0.010319 |
| 237 | COG3934 | Belongs to the glycosyl hydrolase 5 (cellulase A) family | -0.010313 |
| 238 | COG2193 | ferroxidase activity | 0.010305 |
| 239 | COG0668 | transmembrane transport | 0.010281 |
| 240 | COG0602 | queuosine metabolic process | -0.010278 |
| 241 | COG2246 | polysaccharide biosynthetic process | -0.010268 |
| 242 | COG2192 | nodulation | -0.010251 |
| 243 | COG1371 | PFAM Archease protein family (DUF101 UPF0211) | 0.010247 |
| 244 | COG0035 | Catalyzes the conversion of uracil and 5-phospho-alpha- D-ribose 1-diphosphate (PRPP) to UMP and diphosphate | -0.010232 |
| 245 | COG2321 | neutral zinc metallopeptidase | -0.010226 |
| 246 | COG5581 | Acts as a flagellar brake, regulating swimming and swarming in a bis-(3'-5') cyclic diguanylic acid (c-di-GMP)- dependent manner. Binds 1 c-di-GMP dimer per subunit. Increasing levels of c-di-GMP lead to decreased motility | -0.010226 |
| 247 | 33XV7 | Protein of unknown function (DUF3450) | 0.010215 |
| 248 | COG1804 | L-carnitine dehydratase bile acid-inducible protein F | 0.010204 |
| 249 | 345S6 | 0.010198 | |
| 250 | 32TUN | 0.010179 |