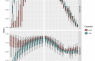
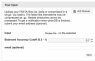
Model weights by orthologous groups
For each model, the rank and weight of this orthologous group are given. These reflect the predictive weight of this protein family for the different traits. Positive weights indicate that the presence of this orthologous group is indicative for the trait. Negative weights indicate that the absence of this orthologous group is indicative for the trait.
| rank in model | enog score | model name | model description |
|---|---|---|---|
| 86 | -0.031712 | for | Organism is producing formic_acid |
| 118 | -0.014628 | T3SS | Organism expresses a Type III secretion system (C) |
| 152 | -0.030941 | etoh | Organism is producing ethanol |
| 186 | -0.011069 | HALO | Organism has a halophilic lifestyle (C) |
| 274 | 0.011393 | PSYCHRO | Organism has a psychrophilic lifestyle (C) |
| 309 | 0.011100 | glc_D | Organism is utilizing D_glucose |
| 319 | -0.009664 | nonfermentative | Organism has a nonfermentative lifestyle |
| 331 | 0.013480 | but | Organism is producing butyric_acid |
| 518 | 0.010160 | Fermentative | Organism has a Fermentative lifestyle |
| 587 | -0.011390 | T6SS | Organism expresses a Type VI secretion system (C) |
| 701 | -0.004539 | AOB | Organism is part of the clade of ammonia-oxidizing bacteria (E) |
| 1283 | -0.007297 | gram_stain | Cells of organism stain Gram-positive (+) or Gram-negative (-) (T) |
| 1378 | -0.004566 | Aerobe | Organism is capable of aerobic respiration (T) |
| 1862 | -0.009682 | ac | Organism is producing acetic_acid |