Quality Analysis of Uploaded Bins (CompleConta)
Completeness/Contamination estimation
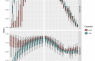
To assess the quality of the bin itself we check the presence or absence of marker genes in the ENOGs returned from the annotation. The 34 universal single copy ENOGs we use were derived from the originally proposed 40 universal protein coding marker gene COGs. The list of those marker genes is used to identify contamination and incompleteness. Completeness and contamination values are used to adjust the expected accuracy of the prediction in later steps.
Taxonomic Placement
The marker genes retrieved also serve as a source to estimate the taxonomic origin of the input bin. We use the bactNOG alignment underlying each ENOG as blastp-database to obtain ncbi-taxids for each sequence that was identified as one of the 34 marker genes. Similar to MEGAN, we select all hits with a score higher than 90% of the best score and perform a last-common-ancestor (LCA) algorithm with a majority rule of again 90%. The LCAs of each sequence are used to calculate an overall LCA for the entire bin, which is returned as the most reliable taxonomic placement.
The scripts used to calculate completeness/contamination and taxonomy can be found on https://github.com/phyden/compleconta