Model Internals
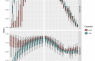
Each PhenDB model is trained on sets of bacterial ENOGs (orthologous groups from EggNOG 4.5), which have or have not been identified in the training genomes. Each ENOG is given a weight, with the magnitude of the weight being the importance of that ENOG for the final prediction. The sign of the weight indicates whether the presence (positive weight) or absence (negative weight) of this ENOG is indicative of the trait.
This table lists the 250 highest-ranking ENOGs of this model.
| rank in model | enog name | enog description | weight in model |
|---|---|---|---|
| 1 | COG0130 | Responsible for synthesis of pseudouridine from uracil- 55 in the psi GC loop of transfer RNAs | 0.174058 |
| 2 | COG1187 | pseudouridine synthase activity | 0.171852 |
| 3 | COG0101 | tRNA pseudouridine synthase activity | 0.171445 |
| 4 | COG0564 | pseudouridine synthase activity | 0.170329 |
| 5 | COG0729 | surface antigen | 0.065328 |
| 6 | COG4977 | sequence-specific DNA binding | 0.064866 |
| 7 | COG0580 | Belongs to the MIP aquaporin (TC 1.A.8) family | 0.063581 |
| 8 | 33AGD | 0.059815 | |
| 9 | 32UTB | 0.059508 | |
| 10 | 30YFD | 0.059495 | |
| 11 | 33Z2K | 0.059494 | |
| 12 | 34AYS | 0.059494 | |
| 13 | COG3226 | Transcriptional regulator | 0.055445 |
| 14 | COG3590 | peptidase | 0.054414 |
| 15 | COG1266 | CAAX protease self-immunity | 0.054301 |
| 16 | COG1376 | ErfK ybiS ycfS ynhG family protein | 0.053093 |
| 17 | COG1012 | belongs to the aldehyde dehydrogenase family | 0.037587 |
| 18 | COG0596 | Alpha beta hydrolase | 0.036515 |
| 19 | COG3239 | Fatty acid desaturase | 0.035138 |
| 20 | COG4638 | Rieske (2fe-2S) | 0.034438 |
| 21 | COG0625 | glutathione transferase activity | 0.033639 |
| 22 | COG5517 | Aromatic-ring-hydroxylating dioxygenase beta subunit | 0.032469 |
| 23 | COG4569 | acetaldehyde dehydrogenase (acetylating) activity | 0.032269 |
| 24 | COG3971 | 2-oxopent-4-enoate hydratase activity | 0.031984 |
| 25 | COG4774 | siderophore transport | 0.030894 |
| 26 | COG3193 | protein, possibly involved in utilization of glycolate and propanediol | 0.030515 |
| 27 | COG0365 | Catalyzes the conversion of acetate into acetyl-CoA (AcCoA), an essential intermediate at the junction of anabolic and catabolic pathways. AcsA undergoes a two-step reaction. In the first half reaction, AcsA combines acetate with ATP to form acetyl-adenylate (AcAMP) intermediate. In the second half reaction, it can then transfer the acetyl group from AcAMP to the sulfhydryl group of CoA, forming the product AcCoA | 0.029773 |
| 28 | COG3917 | 2-hydroxychromene-2-carboxylate isomerase | 0.029734 |
| 29 | COG3347 | Dehydrogenase | 0.027983 |
| 30 | 32TDU | 0.027904 | |
| 31 | COG2146 | nitrite reductase [NAD(P)H] activity | 0.027709 |
| 32 | COG1942 | 4-Oxalocrotonate Tautomerase | 0.026804 |
| 33 | COG0329 | Catalyzes the condensation of (S)-aspartate-beta- semialdehyde (S)-ASA and pyruvate to 4-hydroxy- tetrahydrodipicolinate (HTPA) | 0.026761 |
| 34 | COG2514 | catechol 2,3-dioxygenase activity | 0.026442 |
| 35 | COG1251 | Belongs to the nitrite and sulfite reductase 4Fe-4S domain family | 0.026392 |
| 36 | COG5639 | conserved small protein | 0.026347 |
| 37 | COG1995 | Catalyzes the NAD(P)-dependent oxidation of 4- (phosphohydroxy)-L-threonine (HTP) into 2-amino-3-oxo-4- (phosphohydroxy)butyric acid which spontaneously decarboxylates to form 3-amino-2-oxopropyl phosphate (AHAP) | 0.026307 |
| 38 | COG1062 | S-(hydroxymethyl)glutathione dehydrogenase activity | 0.023312 |
| 39 | COG1961 | COG1961 Site-specific recombinases, DNA invertase Pin homologs | 0.020947 |
| 40 | COG0543 | 2 iron, 2 sulfur cluster binding | 0.020152 |
| 41 | COG1080 | General (non sugar-specific) component of the phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS). This major carbohydrate active-transport system catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. Enzyme I transfers the phosphoryl group from phosphoenolpyruvate (PEP) to the phosphoryl carrier protein (HPr) | 0.019619 |
| 42 | COG2363 | Small membrane protein | 0.018940 |
| 43 | COG0701 | Predicted permease | -0.018143 |
| 44 | COG3604 | Transcriptional regulator | 0.018141 |
| 45 | 2ZAHD | 0.017211 | |
| 46 | 32ZT3 | Thioredoxin domain | -0.017137 |
| 47 | COG0567 | Dehydrogenase, E1 component | 0.017030 |
| 48 | COG2015 | COG2015, Alkyl sulfatase and related hydrolases | 0.016134 |
| 49 | COG2910 | NAD(P)H-binding | 0.016127 |
| 50 | COG3231 | kanamycin kinase activity | 0.015940 |
| 51 | 2ZA7W | Protein of unknown function (DUF1173) | 0.015932 |
| 52 | COG5305 | Membrane | -0.015891 |
| 53 | COG1592 | Rubrerythrin | -0.015792 |
| 54 | 32NDS | Fibronectin type 3 domain | 0.015785 |
| 55 | 30IXR | 0.015656 | |
| 56 | COG0431 | NADPH-dependent FMN reductase | 0.015643 |
| 57 | COG4185 | zeta toxin | 0.015515 |
| 58 | 31UR3 | 0.015496 | |
| 59 | 31H4F | Glyoxalase/Bleomycin resistance protein/Dioxygenase superfamily | 0.015414 |
| 60 | 30WTE | 0.015398 | |
| 61 | COG4624 | Iron only hydrogenase large subunit, C-terminal domain | -0.015362 |
| 62 | 343SX | TrbC/VIRB2 family | 0.015337 |
| 63 | 32T9Z | Protein of unknown function (DUF3169) | 0.015247 |
| 64 | 30S7A | 0.015232 | |
| 65 | COG1331 | Highly conserved protein containing a thioredoxin domain | -0.015220 |
| 66 | 32GX3 | 0.015215 | |
| 67 | 32SUJ | Domain of unknown function (DUF3560) | 0.015167 |
| 68 | COG1070 | Carbohydrate kinase | -0.015088 |
| 69 | 345K5 | 0.015018 | |
| 70 | 2ZCN5 | Type IV secretion system proteins | 0.015006 |
| 71 | COG0613 | PHP domain protein | -0.014987 |
| 72 | 2ZXKF | 0.014947 | |
| 73 | 2ZX78 | 0.014932 | |
| 74 | 2ZXYI | 0.014932 | |
| 75 | 304YF | 0.014932 | |
| 76 | 305D4 | 0.014932 | |
| 77 | 30DSN | 0.014932 | |
| 78 | 31S6W | 0.014932 | |
| 79 | 31SFI | 0.014932 | |
| 80 | 32CEH | 0.014932 | |
| 81 | 32KA0 | 0.014932 | |
| 82 | 32V4T | 0.014932 | |
| 83 | 33AI9 | 0.014932 | |
| 84 | 3481C | 0.014932 | |
| 85 | COG0208 | Provides the precursors necessary for DNA synthesis. Catalyzes the biosynthesis of deoxyribonucleotides from the corresponding ribonucleotides | 0.014918 |
| 86 | COG3329 | Na+-dependent bicarbonate transporter superfamily | 0.014898 |
| 87 | 33DQM | 0.014798 | |
| 88 | COG2963 | transposase activity | 0.014434 |
| 89 | COG2826 | transposase and inactivated derivatives, IS30 family | 0.014308 |
| 90 | COG1834 | amidinotransferase | 0.014290 |
| 91 | COG0475 | glutathione-regulated potassium exporter activity | -0.014223 |
| 92 | COG0411 | ABC transporter | -0.014201 |
| 93 | 32XK3 | Conjugal transfer protein TraD | 0.014195 |
| 94 | COG2169 | Transcriptional regulator | 0.014187 |
| 95 | COG1192 | Involved in chromosome partitioning | 0.014121 |
| 96 | COG3564 | Protein conserved in bacteria | 0.014006 |
| 97 | COG2162 | N-hydroxyarylamine O-acetyltransferase activity | 0.013908 |
| 98 | COG0262 | Key enzyme in folate metabolism. Catalyzes an essential reaction for de novo glycine and purine synthesis, and for DNA precursor synthesis | 0.013821 |
| 99 | COG4232 | protein-disulfide reductase activity | 0.013756 |
| 100 | COG0648 | Endonuclease IV plays a role in DNA repair. It cleaves phosphodiester bonds at apurinic or apyrimidinic sites (AP sites) to produce new 5'-ends that are base-free deoxyribose 5-phosphate residues. It preferentially attacks modified AP sites created by bleomycin and neocarzinostatin | -0.013724 |
| 101 | COG0474 | ATPase, P-type transporting, HAD superfamily, subfamily IC | -0.013636 |
| 102 | COG1072 | (Pantothenic acid kinase)) | 0.013633 |
| 103 | COG1032 | radical SAM domain protein | -0.013574 |
| 104 | COG4637 | Psort location Cytoplasmic, score | -0.013520 |
| 105 | COG4962 | Type ii secretion system protein e | 0.013481 |
| 106 | COG1655 | Protein conserved in bacteria | -0.013435 |
| 107 | COG0459 | protein refolding | 0.013386 |
| 108 | COG1509 | lysine 2,3-aminomutase activity | -0.013368 |
| 109 | COG2006 | 4fe-4S ferredoxin, iron-sulfur binding domain protein | -0.013366 |
| 110 | COG1145 | 4fe-4S ferredoxin, iron-sulfur binding domain protein | -0.013316 |
| 111 | COG3449 | Inhibits the supercoiling activity of DNA gyrase. Acts by inhibiting DNA gyrase at an early step, prior to (or at the step of) binding of DNA by the gyrase. It protects cells against toxins that target DNA gyrase, by inhibiting activity of these toxins and reducing the formation of lethal double-strand breaks in the cell | -0.013216 |
| 112 | COG0508 | dehydrogenase complex catalyzes the overall conversion of | 0.013033 |
| 113 | COG1350 | The beta subunit is responsible for the synthesis of L- tryptophan from indole and L-serine | -0.012973 |
| 114 | COG0410 | ABC transporter | -0.012963 |
| 115 | 345M7 | C4-type zinc ribbon domain | -0.012951 |
| 116 | COG2346 | COG2346, Truncated hemoglobins | 0.012922 |
| 117 | COG1151 | hydroxylamine reductase activity | -0.012918 |
| 118 | COG1550 | Protein conserved in bacteria | -0.012861 |
| 119 | COG0684 | Catalyzes the aldol cleavage of 4-hydroxy-4-methyl-2- oxoglutarate (HMG) into 2 molecules of pyruvate. Also contains a secondary oxaloacetate (OAA) decarboxylase activity due to the common pyruvate enolate transition state formed following C-C bond cleavage in the retro-aldol and decarboxylation reactions | 0.012662 |
| 120 | COG4177 | L-phenylalanine transmembrane transporter activity | -0.012644 |
| 121 | COG0674 | pyruvate-flavodoxin oxidoreductase activity | -0.012637 |
| 122 | COG0420 | 3'-5' exonuclease activity | 0.012606 |
| 123 | COG2210 | Belongs to the sulfur carrier protein TusA family | -0.012517 |
| 124 | COG3877 | Protein conserved in bacteria | -0.012483 |
| 125 | COG0234 | Binds to Cpn60 in the presence of Mg-ATP and suppresses the ATPase activity of the latter | 0.012439 |
| 126 | COG3702 | type iv secretory pathway | 0.012381 |
| 127 | COG2048 | Heterodisulfide reductase, subunit B | -0.012356 |
| 128 | COG3664 | PFAM glycoside hydrolase family 39 | -0.012313 |
| 129 | COG1278 | Cold shock | 0.012287 |
| 130 | COG1014 | Oxidoreductase required for the transfer of electrons from pyruvate to flavodoxin | -0.012264 |
| 131 | COG2842 | Transposition protein | -0.012205 |
| 132 | COG1304 | FMN binding | 0.012172 |
| 133 | COG0559 | Belongs to the binding-protein-dependent transport system permease family | -0.012105 |
| 134 | COG1525 | nuclease | 0.012057 |
| 135 | COG3182 | Iron-regulated membrane protein | 0.011941 |
| 136 | COG0523 | cobalamin synthesis protein | 0.011940 |
| 137 | COG3504 | Conjugal transfer protein | 0.011851 |
| 138 | COG1342 | Belongs to the UPF0251 family | -0.011781 |
| 139 | COG1668 | transmembrane transport | -0.011684 |
| 140 | COG3033 | Beta-eliminating lyase | -0.011682 |
| 141 | COG2030 | dehydratase | 0.011646 |
| 142 | COG1647 | Serine aminopeptidase, S33 | 0.011625 |
| 143 | COG1328 | CTP reductase activity | -0.011568 |
| 144 | COG2605 | GHMP kinase | -0.011558 |
| 145 | COG1351 | Catalyzes the reductive methylation of 2'-deoxyuridine- 5'-monophosphate (dUMP) to 2'-deoxythymidine-5'-monophosphate (dTMP) while utilizing 5,10-methylenetetrahydrofolate (mTHF) as the methyl donor, and NAD(P)H and FADH(2) as the reductant | -0.011548 |
| 146 | COG2221 | Nitrite and sulphite reductase 4Fe-4S | -0.011542 |
| 147 | COG1324 | tolerance protein | 0.011498 |
| 148 | COG4656 | Part of a membrane complex involved in electron transport | -0.011465 |
| 149 | COG3736 | Type IV secretory pathway, component VirB8 | 0.011413 |
| 150 | COG0604 | alcohol dehydrogenase | 0.011355 |
| 151 | COG3620 | sequence-specific DNA binding | 0.011199 |
| 152 | COG0279 | D-glycero-D-manno-heptose 7-phosphate biosynthetic process | -0.011195 |
| 153 | COG2050 | protein possibly involved in aromatic compounds catabolism | 0.011192 |
| 154 | COG3808 | pump that utilizes the energy of pyrophosphate hydrolysis as the driving force for | -0.011153 |
| 155 | COG3505 | Type IV secretory pathway VirD4 | 0.011124 |
| 156 | COG0316 | Belongs to the HesB IscA family | 0.011042 |
| 157 | COG4559 | Part of the ABC transporter complex HmuTUV involved in hemin import. Responsible for energy coupling to the transport system | -0.011029 |
| 158 | COG0496 | Nucleotidase that shows phosphatase activity on nucleoside 5'-monophosphates | -0.011015 |
| 159 | COG1752 | Esterase of the alpha-beta hydrolase superfamily | -0.011008 |
| 160 | COG0432 | Pfam Uncharacterised protein family UPF0047 | -0.010880 |
| 161 | COG1030 | water channel activity | -0.010857 |
| 162 | COG1908 | Methyl-viologen-reducing hydrogenase, delta subunit | -0.010836 |
| 163 | COG1245 | 4Fe-4S binding domain | -0.010834 |
| 164 | COG2303 | Involved in the biosynthesis of the osmoprotectant glycine betaine. Catalyzes the oxidation of choline to betaine aldehyde and betaine aldehyde to glycine betaine at the same rate | 0.010833 |
| 165 | COG1003 | The glycine cleavage system catalyzes the degradation of glycine. The P protein binds the alpha-amino group of glycine through its pyridoxal phosphate cofactor | -0.010821 |
| 166 | COG4733 | cellulase activity | -0.010781 |
| 167 | COG1013 | pyruvate-flavodoxin oxidoreductase activity | -0.010734 |
| 168 | COG4623 | carbon-oxygen lyase activity, acting on polysaccharides | 0.010646 |
| 169 | COG0026 | Catalyzes the ATP-dependent conversion of 5- aminoimidazole ribonucleotide (AIR) and HCO(3)(-) to N5- carboxyaminoimidazole ribonucleotide (N5-CAIR) | 0.010644 |
| 170 | COG3704 | protein secretion by the type IV secretion system | 0.010610 |
| 171 | COG0632 | four-way junction helicase activity | 0.010537 |
| 172 | COG2348 | transferase activity, transferring amino-acyl groups | -0.010406 |
| 173 | COG1089 | Catalyzes the conversion of GDP-D-mannose to GDP-4- dehydro-6-deoxy-D-mannose | -0.010377 |
| 174 | COG1313 | radical SAM domain protein | -0.010363 |
| 175 | COG1023 | phosphogluconate dehydrogenase (decarboxylating) activity | -0.010347 |
| 176 | COG2089 | acid synthase | -0.010337 |
| 177 | COG5006 | permease, DMT superfamily | 0.010291 |
| 178 | COG5504 | Zn-dependent protease | 0.010262 |
| 179 | COG1083 | cytidylyl-transferase | -0.010178 |
| 180 | COG1139 | Iron-sulfur cluster binding protein | -0.010169 |
| 181 | COG0757 | Catalyzes a trans-dehydration via an enolate intermediate | -0.010127 |
| 182 | COG1876 | peptidase activity | 0.010098 |
| 183 | COG5512 | Zn-ribbon-containing possibly RNA-binding protein and truncated derivatives | -0.010093 |
| 184 | COG4119 | NUDIX hydrolase | -0.010026 |
| 185 | COG3144 | bacterial-type flagellum assembly | -0.009970 |
| 186 | COG0683 | leucine binding | -0.009970 |
| 187 | COG4108 | Increases the formation of ribosomal termination complexes and stimulates activities of RF-1 and RF-2. It binds guanine nucleotides and has strong preference for UGA stop codons. It may interact directly with the ribosome. The stimulation of RF- 1 and RF-2 is significantly reduced by GTP and GDP, but not by GMP | 0.009953 |
| 188 | COG1449 | Belongs to the glycosyl hydrolase 57 family | -0.009938 |
| 189 | COG3262 | NADH dehydrogenase (ubiquinone), 30 kDa subunit | -0.009922 |
| 190 | COG1695 | Transcriptional regulator | 0.009905 |
| 191 | 345AV | Cyclic nucleotide-monophosphate binding domain | -0.009897 |
| 192 | COG3576 | pyridoxamine 5-phosphate | -0.009879 |
| 193 | COG2844 | Modifies, by uridylylation and deuridylylation, the PII regulatory proteins (GlnB and homologs), in response to the nitrogen status of the cell that GlnD senses through the glutamine level. Under low glutamine levels, catalyzes the conversion of the PII proteins and UTP to PII-UMP and PPi, while under higher glutamine levels, GlnD hydrolyzes PII-UMP to PII and UMP (deuridylylation). Thus, controls uridylylation state and activity of the PII proteins, and plays an important role in the regulation of nitrogen | -0.009870 |
| 194 | COG1294 | oxidase, subunit | 0.009855 |
| 195 | COG0122 | 3-methyladenine DNA glycosylase 8-oxoguanine DNA glycosylase | 0.009834 |
| 196 | COG1666 | GTP binding | 0.009818 |
| 197 | COG0276 | Catalyzes the ferrous insertion into protoporphyrin IX | 0.009775 |
| 198 | COG1180 | Activation of pyruvate formate-lyase under anaerobic conditions by generation of an organic free radical, using S- adenosylmethionine and reduced flavodoxin as cosubstrates to produce 5'-deoxy-adenosine | -0.009772 |
| 199 | COG4843 | Uncharacterized protein conserved in bacteria (DUF2179) | -0.009767 |
| 200 | COG4097 | nitric oxide dioxygenase activity | 0.009750 |
| 201 | COG1492 | cobalamin metabolic process | -0.009747 |
| 202 | COG3426 | butyrate kinase activity | -0.009729 |
| 203 | COG0509 | glycine decarboxylation via glycine cleavage system | -0.009677 |
| 204 | COG2948 | multi-organism process | 0.009659 |
| 205 | COG1764 | response to oxidative stress | 0.009650 |
| 206 | COG3393 | -acetyltransferase | 0.009634 |
| 207 | COG1905 | PFAM NADH dehydrogenase (ubiquinone) 24 kDa subunit | -0.009606 |
| 208 | COG3635 | 2,3-bisphosphoglycerate-independent phosphoglycerate mutase activity | -0.009603 |
| 209 | COG1686 | Belongs to the peptidase S11 family | 0.009587 |
| 210 | COG3655 | Transcriptional regulator | 0.009578 |
| 211 | COG3221 | ABC-type phosphate phosphonate transport system periplasmic component | -0.009572 |
| 212 | COG1514 | Hydrolyzes RNA 2',3'-cyclic phosphodiester to an RNA 2'- phosphomonoester | -0.009498 |
| 213 | COG0803 | Belongs to the bacterial solute-binding protein 9 family | -0.009485 |
| 214 | COG3961 | Belongs to the TPP enzyme family | 0.009463 |
| 215 | COG2309 | aminopeptidase activity | -0.009460 |
| 216 | COG3853 | Toxic anion resistance protein (TelA) | 0.009454 |
| 217 | COG1002 | Type II restriction enzyme, methylase subunits | -0.009441 |
| 218 | COG1516 | flagellar protein fliS | -0.009433 |
| 219 | COG3324 | translation initiation factor activity | 0.009396 |
| 220 | COG3284 | Transcriptional regulator | 0.009387 |
| 221 | COG0109 | protoheme IX farnesyltransferase activity | 0.009362 |
| 222 | COG0827 | DNA restriction-modification system | 0.009338 |
| 223 | COG0040 | Catalyzes the condensation of ATP and 5-phosphoribose 1- diphosphate to form N'-(5'-phosphoribosyl)-ATP (PR-ATP). Has a crucial role in the pathway because the rate of histidine biosynthesis seems to be controlled primarily by regulation of hisG enzymatic activity | -0.009336 |
| 224 | COG3384 | dioxygenase | 0.009328 |
| 225 | COG2072 | flavoprotein involved in K transport | 0.009266 |
| 226 | COG3024 | Inhibits all the catalytic activities of DNA gyrase by preventing its interaction with DNA. Acts by binding directly to the C-terminal domain of GyrB, which probably disrupts DNA binding by the gyrase | 0.009260 |
| 227 | COG2185 | Catalyzes the reversible interconversion of isobutyryl- CoA and n-butyryl-CoA, using radical chemistry. Also exhibits GTPase activity, associated with its G-protein domain (MeaI) that functions as a chaperone that assists cofactor delivery and proper holo-enzyme assembly | -0.009258 |
| 228 | COG0650 | Formate hydrogenlyase subunit 4 | -0.009248 |
| 229 | 33YJY | -0.009234 | |
| 230 | COG2609 | component of the pyruvate dehydrogenase (PDH) complex, that catalyzes the overall conversion of pyruvate to acetyl-CoA and CO(2) | 0.009227 |
| 231 | COG3973 | AAA domain | 0.009217 |
| 232 | COG4798 | methyltransferase | -0.009213 |
| 233 | COG3303 | Catalyzes the reduction of nitrite to ammonia, consuming six electrons in the process | -0.009207 |
| 234 | COG1150 | heterodisulfide reductase subunit C | -0.009206 |
| 235 | COG2828 | Protein conserved in bacteria | 0.009204 |
| 236 | COG0680 | Initiates the rapid degradation of small, acid-soluble proteins during spore germination | -0.009193 |
| 237 | COG1226 | (belongs to the monovalent cation proton antiporter 2 (CPA2) transporter (TC 2.A.37) family) | 0.009182 |
| 238 | COG0583 | DNA-binding transcription factor activity | 0.009166 |
| 239 | COG0342 | Part of the Sec protein translocase complex. Interacts with the SecYEG preprotein conducting channel. SecDF uses the proton motive force (PMF) to complete protein translocation after the ATP-dependent function of SecA | -0.009152 |
| 240 | COG4584 | PFAM Integrase catalytic | -0.009131 |
| 241 | COG1566 | PFAM secretion protein HlyD family protein | -0.009105 |
| 242 | COG0106 | 1-(5-phosphoribosyl)-5- 5-phosphoribosylamino)methylideneamino imidazole-4-carboxamide isomerase | -0.009098 |
| 243 | COG0207 | Catalyzes the reductive methylation of 2'-deoxyuridine- 5'-monophosphate (dUMP) to 2'-deoxythymidine-5'-monophosphate (dTMP) while utilizing 5,10-methylenetetrahydrofolate (mTHF) as the methyl donor and reductant in the reaction, yielding dihydrofolate (DHF) as a by-product. This enzymatic reaction provides an intracellular de novo source of dTMP, an essential precursor for DNA biosynthesis | 0.009082 |
| 244 | COG0753 | catalase activity | 0.009082 |
| 245 | COG1237 | beta-lactamase domain protein | -0.009077 |
| 246 | 3482A | -0.009073 | |
| 247 | COG1703 | isobutyryl-CoA mutase activity | -0.009057 |
| 248 | COG3540 | Alkaline phosphatase | 0.009043 |
| 249 | COG1454 | alcohol dehydrogenase | -0.009024 |
| 250 | COG0692 | Excises uracil residues from the DNA which can arise as a result of misincorporation of dUMP residues by DNA polymerase or due to deamination of cytosine | 0.009015 |