Model Internals
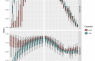
Each PhenDB model is trained on sets of bacterial ENOGs (orthologous groups from EggNOG 4.5), which have or have not been identified in the training genomes. Each ENOG is given a weight, with the magnitude of the weight being the importance of that ENOG for the final prediction. The sign of the weight indicates whether the presence (positive weight) or absence (negative weight) of this ENOG is indicative of the trait.
This table lists the 250 highest-ranking ENOGs of this model.
| rank in model | enog name | enog description | weight in model |
|---|---|---|---|
| 1 | COG1063 | alcohol dehydrogenase | -0.060357 |
| 2 | COG3711 | transcriptional antiterminator | 0.055179 |
| 3 | COG1757 | Na H antiporter | -0.052681 |
| 4 | COG3775 | system Galactitol-specific IIC component | 0.050069 |
| 5 | COG1322 | Protein conserved in bacteria | 0.049710 |
| 6 | COG3633 | Involved in the import of serine and threonine into the cell, with the concomitant import of sodium (symport system) | 0.049460 |
| 7 | COG5586 | Nucleotidyl transferase AbiEii toxin, Type IV TA system | -0.048192 |
| 8 | 31EGY | 0.047542 | |
| 9 | COG0562 | UDP-galactopyranose mutase | -0.047273 |
| 10 | COG2378 | regulation of single-species biofilm formation | -0.045144 |
| 11 | COG2116 | formate transmembrane transporter activity | 0.044467 |
| 12 | COG3465 | dGTPase activity | -0.044063 |
| 13 | COG0655 | NAD(P)H dehydrogenase (quinone) activity | 0.044008 |
| 14 | 2Z8AD | Carbohydrate-binding domain-containing protein Cthe_2159 | 0.043516 |
| 15 | 2ZSRR | Psort location Cytoplasmic, score 8.87 | 0.042664 |
| 16 | COG3560 | Nitroreductase | -0.042487 |
| 17 | COG4587 | transport system, permease component | 0.042293 |
| 18 | COG1794 | racemase activity, acting on amino acids and derivatives | -0.042152 |
| 19 | COG3247 | response to pH | 0.041678 |
| 20 | COG2818 | Glycosylase | -0.041022 |
| 21 | COG1479 | Protein of unknown function DUF262 | 0.041021 |
| 22 | COG1619 | proteins homologs of microcin C7 resistance protein MccF | 0.040353 |
| 23 | COG2827 | Endonuclease containing a URI domain | -0.039899 |
| 24 | 33P14 | FIVAR domain | 0.039659 |
| 25 | COG4245 | von Willebrand factor, type A | 0.038789 |
| 26 | COG5360 | Heparinase II/III-like protein | 0.038619 |
| 27 | COG4225 | unsaturated rhamnogalacturonyl hydrolase activity | 0.038504 |
| 28 | COG3513 | CRISPR (clustered regularly interspaced short palindromic repeat) is an adaptive immune system that provides protection against mobile genetic elements (viruses, transposable elements and conjugative plasmids). CRISPR clusters contain spacers, sequences complementary to antecedent mobile elements, and target invading nucleic acids. CRISPR clusters are transcribed and processed into CRISPR RNA (crRNA). In type II CRISPR systems correct processing of pre-crRNA requires a trans-encoded small RNA (tracrRNA), endogenous ribonuclease 3 (rnc) and this protein. The tracrRNA serves as a guide for ribonuclease 3-aided processing of pre-crRNA. Subsequently Cas9 crRNA tracrRNA endonucleolytically cleaves linear or circular dsDNA target complementary to the spacer | -0.038422 |
| 29 | COG4559 | Part of the ABC transporter complex HmuTUV involved in hemin import. Responsible for energy coupling to the transport system | -0.038250 |
| 30 | COG0615 | ADP-L-glycero-beta-D-manno-heptose biosynthetic process | -0.038166 |
| 31 | COG3589 | Outer surface protein | 0.037908 |
| 32 | COG3191 | aminopeptidase activity | -0.037898 |
| 33 | COG3263 | regulation of cellular component size | -0.037785 |
| 34 | COG1233 | COG1233 Phytoene dehydrogenase and related | -0.037550 |
| 35 | COG0846 | NAD-dependent lysine deacetylase and desuccinylase that specifically removes acetyl and succinyl groups on target proteins. Modulates the activities of several proteins which are inactive in their acylated form | -0.037416 |
| 36 | COG1053 | succinate dehydrogenase | -0.037108 |
| 37 | COG3221 | ABC-type phosphate phosphonate transport system periplasmic component | -0.036944 |
| 38 | 32TBR | Protein of unknown function (DUF2974) | 0.036866 |
| 39 | COG3275 | phosphorelay sensor kinase activity | 0.036770 |
| 40 | 2ZIYW | Psort location Cytoplasmic, score | 0.036712 |
| 41 | COG0584 | glycerophosphoryl diester phosphodiesterase | -0.036173 |
| 42 | COG4642 | MORN repeat | 0.036079 |
| 43 | COG3408 | Glycogen debranching enzyme | 0.035924 |
| 44 | COG4095 | Psort location CytoplasmicMembrane, score | -0.035659 |
| 45 | COG5340 | Psort location Cytoplasmic, score | -0.035439 |
| 46 | COG1680 | COG1680 Beta-lactamase class C and other penicillin binding | -0.035427 |
| 47 | COG1971 | Probably functions as a manganese efflux pump | 0.035420 |
| 48 | COG1882 | Formate acetyltransferase | 0.035199 |
| 49 | COG5499 | transcription regulator containing HTH domain | 0.034952 |
| 50 | 2Z8ZV | ABC-2 family transporter protein | -0.034918 |
| 51 | COG4720 | Psort location CytoplasmicMembrane, score | 0.034850 |
| 52 | COG0785 | Cytochrome C biogenesis protein | -0.034532 |
| 53 | COG0715 | COG0715 ABC-type nitrate sulfonate bicarbonate transport systems periplasmic components | 0.034479 |
| 54 | 32XKY | ECF transporter, substrate-specific component | -0.034442 |
| 55 | COG0471 | metal ion transport | -0.034377 |
| 56 | COG0794 | Belongs to the SIS family. GutQ KpsF subfamily | -0.034274 |
| 57 | COG5485 | Ester cyclase | 0.033959 |
| 58 | COG1748 | Saccharopine dehydrogenase | 0.033923 |
| 59 | COG3833 | ABC-type maltose transport systems, permease component | 0.033795 |
| 60 | COG1916 | peptidoglycan binding | 0.033773 |
| 61 | COG1080 | General (non sugar-specific) component of the phosphoenolpyruvate-dependent sugar phosphotransferase system (sugar PTS). This major carbohydrate active-transport system catalyzes the phosphorylation of incoming sugar substrates concomitantly with their translocation across the cell membrane. Enzyme I transfers the phosphoryl group from phosphoenolpyruvate (PEP) to the phosphoryl carrier protein (HPr) | 0.033403 |
| 62 | COG0390 | transport system, permease component | 0.033318 |
| 63 | 33CG9 | GDSL-like Lipase/Acylhydrolase | 0.033314 |
| 64 | COG4915 | 5-bromo-4-chloroindolyl phosphate hydrolysis protein | -0.033277 |
| 65 | COG0801 | 2-amino-4-hydroxy-6-hydroxymethyldihydropteridine pyrophosphokinase | 0.033156 |
| 66 | COG1180 | Activation of pyruvate formate-lyase under anaerobic conditions by generation of an organic free radical, using S- adenosylmethionine and reduced flavodoxin as cosubstrates to produce 5'-deoxy-adenosine | 0.033010 |
| 67 | COG3459 | Glycosyltransferase 36 associated | 0.032881 |
| 68 | COG3967 | Belongs to the short-chain dehydrogenases reductases (SDR) family | -0.032862 |
| 69 | 32RYJ | Domain of unknown function (DUF4186) | 0.032846 |
| 70 | COG0210 | DNA helicase | 0.032799 |
| 71 | COG3618 | amidohydrolase | 0.032680 |
| 72 | COG1429 | ligase activity, forming nitrogen-metal bonds | -0.032669 |
| 73 | COG1669 | hydrolase activity, acting on ester bonds | 0.032624 |
| 74 | COG2129 | metallophosphoesterase | -0.032614 |
| 75 | COG0421 | Catalyzes the irreversible transfer of a propylamine group from the amino donor S-adenosylmethioninamine (decarboxy- AdoMet) to putrescine (1,4-diaminobutane) to yield spermidine | 0.032442 |
| 76 | COG1989 | Cleaves type-4 fimbrial leader sequence and methylates the N-terminal (generally Phe) residue | -0.032432 |
| 77 | COG1917 | Cupin 2, conserved barrel domain protein | -0.032398 |
| 78 | COG1278 | Cold shock | -0.032188 |
| 79 | COG3257 | ureidoglycine aminohydrolase activity | 0.031884 |
| 80 | COG4221 | oxidoreductase activity | 0.031883 |
| 81 | 303R9 | -0.031817 | |
| 82 | COG4824 | toxin secretion phage lysis holin | -0.031810 |
| 83 | COG1508 | bacterial-type RNA polymerase transcriptional activator activity, sequence-specific DNA binding | -0.031782 |
| 84 | COG1442 | glycosyl transferase family 8 | -0.031767 |
| 85 | COG1074 | ATP-dependent DNA helicase activity | -0.031763 |
| 86 | COG2509 | 5-formyltetrahydrofolate cyclo-ligase activity | -0.031712 |
| 87 | COG4284 | Utp--glucose-1-phosphate uridylyltransferase | 0.031703 |
| 88 | COG3393 | -acetyltransferase | -0.031669 |
| 89 | 338B2 | 0.031639 | |
| 90 | COG3639 | organic phosphonate transmembrane transporter activity | -0.031534 |
| 91 | 32U4Y | helix_turn_helix, Lux Regulon | 0.031507 |
| 92 | COG0063 | ADP-dependent NAD(P)H-hydrate dehydratase activity | -0.031391 |
| 93 | COG1652 | LysM domain | -0.031370 |
| 94 | COG3830 | Belongs to the UPF0237 family | 0.031363 |
| 95 | 342R0 | Trypsin-like peptidase domain | -0.031275 |
| 96 | COG5337 | Spore coat protein CotH | 0.031149 |
| 97 | COG2113 | Glycine betaine | 0.031142 |
| 98 | COG5617 | Psort location CytoplasmicMembrane, score | -0.031095 |
| 99 | COG1636 | Catalyzes the conversion of epoxyqueuosine (oQ) to queuosine (Q), which is a hypermodified base found in the wobble positions of tRNA(Asp), tRNA(Asn), tRNA(His) and tRNA(Tyr) | -0.030956 |
| 100 | COG3757 | hydrolase, family 25 | 0.030916 |
| 101 | COG1922 | lipopolysaccharide N-acetylmannosaminouronosyltransferase activity | -0.030893 |
| 102 | COG1062 | S-(hydroxymethyl)glutathione dehydrogenase activity | 0.030820 |
| 103 | COG2964 | Protein conserved in bacteria | 0.030782 |
| 104 | COG0393 | Putative heavy-metal-binding | 0.030778 |
| 105 | COG1741 | Belongs to the pirin family | -0.030770 |
| 106 | COG2271 | Major facilitator Superfamily | -0.030745 |
| 107 | 2Z7TS | Domain of unknown function (DUF4317) | 0.030726 |
| 108 | COG2452 | N-4 methylation of cytosine | -0.030719 |
| 109 | COG5566 | PFAM Mor transcription activator | -0.030654 |
| 110 | COG4886 | Leucine-rich repeat (LRR) protein | 0.030635 |
| 111 | COG3037 | sugar-specific permease SgaT UlaA | 0.030614 |
| 112 | 2ZYY6 | Bacterial regulatory proteins, tetR family | -0.030606 |
| 113 | COG3598 | Psort location Cytoplasmic, score | 0.030530 |
| 114 | COG3740 | Phage prohead protease, HK97 family | -0.030506 |
| 115 | COG3290 | protein histidine kinase activity | 0.030438 |
| 116 | 32S7G | Excisionase | 0.030372 |
| 117 | COG1477 | protein flavinylation | -0.030354 |
| 118 | COG0235 | Class ii aldolase | 0.030337 |
| 119 | COG1827 | regulation of RNA biosynthetic process | 0.030232 |
| 120 | COG1038 | Catalyzes a 2-step reaction, involving the ATP-dependent carboxylation of the covalently attached biotin in the first step and the transfer of the carboxyl group to pyruvate in the second | -0.030168 |
| 121 | COG3707 | response regulator | 0.030150 |
| 122 | COG2845 | Protein of unknown function (DUF459) | -0.030115 |
| 123 | 30TEW | Ribosomal protein L33 | 0.030073 |
| 124 | 32AMX | -0.029744 | |
| 125 | COG4694 | Protein conserved in bacteria | -0.029714 |
| 126 | COG4235 | cytochrome complex assembly | 0.029650 |
| 127 | COG0351 | phosphomethylpyrimidine kinase | 0.029649 |
| 128 | COG1416 | DNA-binding transcription factor activity | 0.029571 |
| 129 | COG2227 | 3-demethylubiquinone-9 3-O-methyltransferase activity | -0.029443 |
| 130 | COG0557 | exoribonuclease II activity | -0.029386 |
| 131 | COG5016 | pyruvate | 0.029155 |
| 132 | COG1541 | Catalyzes the activation of phenylacetic acid (PA) to phenylacetyl-CoA (PA-CoA) | 0.029141 |
| 133 | COG0174 | glutamine synthetase | 0.029063 |
| 134 | COG2030 | dehydratase | -0.028997 |
| 135 | COG3878 | Protein conserved in bacteria | -0.028943 |
| 136 | COG0395 | ABC-type sugar transport system, permease component | 0.028921 |
| 137 | COG4302 | ethanolamine ammonia-lyase activity | 0.028909 |
| 138 | COG2876 | 3-deoxy-7-phosphoheptulonate synthase activity | -0.028895 |
| 139 | COG3304 | Inner membrane component domain | 0.028875 |
| 140 | COG2138 | Cobalamin (vitamin B12) biosynthesis CbiX protein | -0.028794 |
| 141 | COG3272 | Protein of unknown function (DUF1722) | 0.028790 |
| 142 | COG1380 | Effector of murein hydrolase LrgA | -0.028789 |
| 143 | 312PY | Cro/C1-type HTH DNA-binding domain | -0.028787 |
| 144 | COG1864 | DNA RNA non-specific endonuclease | -0.028663 |
| 145 | COG2862 | Uncharacterized protein family, UPF0114 | -0.028647 |
| 146 | COG1328 | CTP reductase activity | -0.028370 |
| 147 | COG3576 | pyridoxamine 5-phosphate | 0.028252 |
| 148 | COG1132 | (ABC) transporter | -0.028174 |
| 149 | COG0721 | Allows the formation of correctly charged Asn-tRNA(Asn) or Gln-tRNA(Gln) through the transamidation of misacylated Asp- tRNA(Asn) or Glu-tRNA(Gln) in organisms which lack either or both of asparaginyl-tRNA or glutaminyl-tRNA synthetases. The reaction takes place in the presence of glutamine and ATP through an activated phospho-Asp-tRNA(Asn) or phospho-Glu-tRNA(Gln) | -0.028164 |
| 150 | COG3467 | pyridoxamine 5'-phosphate | -0.028058 |
| 151 | COG2524 | Domain in cystathionine beta-synthase and other proteins. | -0.028051 |
| 152 | COG3226 | Transcriptional regulator | 0.028032 |
| 153 | 340N5 | 0.027998 | |
| 154 | COG1041 | phosphinothricin N-acetyltransferase activity | 0.027950 |
| 155 | COG0616 | signal peptide processing | -0.027883 |
| 156 | COG2972 | Histidine kinase | 0.027879 |
| 157 | COG2235 | arginine | -0.027870 |
| 158 | COG1525 | nuclease | -0.027866 |
| 159 | COG0007 | Multifunctional enzyme that catalyzes the SAM-dependent methylations of uroporphyrinogen III at position C-2 and C-7 to form precorrin-2 via precorrin-1. Then it catalyzes the NAD- dependent ring dehydrogenation of precorrin-2 to yield sirohydrochlorin. Finally, it catalyzes the ferrochelation of sirohydrochlorin to yield siroheme | -0.027863 |
| 160 | 33A5I | Psort location CytoplasmicMembrane, score | 0.027858 |
| 161 | COG4894 | glucomannan catabolic process | 0.027857 |
| 162 | 2ZGPA | PspA/IM30 family | 0.027826 |
| 163 | COG0176 | Transaldolase is important for the balance of metabolites in the pentose-phosphate pathway | 0.027810 |
| 164 | COG4702 | Belongs to the UPF0303 family | 0.027766 |
| 165 | 33CTH | MORN repeat variant | -0.027761 |
| 166 | 33BGI | Parallel beta-helix repeats | 0.027747 |
| 167 | 30GAQ | GDSL-like Lipase/Acylhydrolase family | -0.027719 |
| 168 | COG1305 | Transglutaminase-like | -0.027627 |
| 169 | COG0154 | amidase activity | -0.027569 |
| 170 | COG0741 | lytic transglycosylase activity | 0.027528 |
| 171 | COG2267 | carboxylic ester hydrolase activity | 0.027512 |
| 172 | COG3328 | transposase activity | -0.027492 |
| 173 | COG4187 | Peptidase family M20/M25/M40 | -0.027464 |
| 174 | COG5546 | COG5546 Small integral membrane protein | 0.027462 |
| 175 | COG3433 | isochorismatase activity | 0.027427 |
| 176 | 33E24 | Helix-turn-helix XRE-family like proteins | -0.027399 |
| 177 | COG2364 | Membrane | -0.027385 |
| 178 | COG4626 | Terminase | -0.027352 |
| 179 | COG3525 | Glycosyl hydrolase, family 20, catalytic domain | 0.027196 |
| 180 | COG4965 | Type ii secretion system | 0.027144 |
| 181 | 33PG8 | Psort location Cytoplasmic, score 8.96 | -0.027116 |
| 182 | COG2216 | Part of the high-affinity ATP-driven potassium transport (or Kdp) system, which catalyzes the hydrolysis of ATP coupled with the electrogenic transport of potassium into the cytoplasm. This subunit is responsible for energy coupling to the transport system | -0.027103 |
| 183 | COG0296 | Catalyzes the formation of the alpha-1,6-glucosidic linkages in glycogen by scission of a 1,4-alpha-linked oligosaccharide from growing alpha-1,4-glucan chains and the subsequent attachment of the oligosaccharide to the alpha-1,6 position | 0.027080 |
| 184 | 32W9S | -0.027009 | |
| 185 | COG1182 | Catalyzes the reductive cleavage of azo bond in aromatic azo compounds to the corresponding amines. Requires NADH, but not NADPH, as an electron donor for its activity | -0.027006 |
| 186 | COG3412 | dihydroxyacetone kinase, phosphotransfer subunit | 0.026983 |
| 187 | COG1879 | ABC-type sugar transport system periplasmic component | -0.026947 |
| 188 | COG1510 | regulation of RNA biosynthetic process | 0.026942 |
| 189 | COG0402 | S-adenosylhomocysteine deaminase activity | -0.026928 |
| 190 | COG0526 | COG0526, thiol-disulfide isomerase and thioredoxins | 0.026912 |
| 191 | 2ZC3Y | family 11 | -0.026864 |
| 192 | COG2382 | esterase | 0.026828 |
| 193 | 31FR9 | MarR family | 0.026794 |
| 194 | COG1286 | toxin biosynthetic process | -0.026763 |
| 195 | COG1329 | Transcriptional regulator | 0.026745 |
| 196 | COG1297 | OPT oligopeptide transporter protein | -0.026734 |
| 197 | COG1142 | 4fe-4S ferredoxin, iron-sulfur binding domain protein | 0.026708 |
| 198 | COG2896 | Catalyzes the cyclization of GTP to (8S)-3',8-cyclo-7,8- dihydroguanosine 5'-triphosphate | -0.026681 |
| 199 | COG0420 | 3'-5' exonuclease activity | -0.026658 |
| 200 | COG5017 | Glycosyltransferase family 28 C-terminal domain | -0.026654 |
| 201 | COG2431 | L-lysine efflux transmembrane transporter activity | -0.026622 |
| 202 | COG4927 | Acyl-coenzyme A 6-aminopenicillanic acid acyl-transferase | -0.026607 |
| 203 | 342VC | Cyclic nucleotide-monophosphate binding domain | -0.026566 |
| 204 | COG0161 | adenosylmethionine-8-amino-7-oxononanoate transaminase activity | -0.026556 |
| 205 | COG0672 | )-iron permease | -0.026550 |
| 206 | COG1023 | phosphogluconate dehydrogenase (decarboxylating) activity | 0.026532 |
| 207 | COG1116 | anion transmembrane transporter activity | 0.026515 |
| 208 | 2ZD1U | 0.026495 | |
| 209 | 32RP2 | Psort location CytoplasmicMembrane, score | 0.026482 |
| 210 | COG3254 | rhamnose metabolic process | 0.026439 |
| 211 | COG5602 | Belongs to the OprB family | 0.026408 |
| 212 | COG3728 | DNA packaging | -0.026392 |
| 213 | COG3731 | PTS system glucitol sorbitol-specific IIA component | -0.026378 |
| 214 | COG1473 | amidohydrolase | -0.026331 |
| 215 | 33REM | Sigma-70 region 2 | -0.026318 |
| 216 | COG2160 | L-arabinose isomerase activity | 0.026275 |
| 217 | COG0064 | Allows the formation of correctly charged Asn-tRNA(Asn) or Gln-tRNA(Gln) through the transamidation of misacylated Asp- tRNA(Asn) or Glu-tRNA(Gln) in organisms which lack either or both of asparaginyl-tRNA or glutaminyl-tRNA synthetases. The reaction takes place in the presence of glutamine and ATP through an activated phospho-Asp-tRNA(Asn) or phospho-Glu-tRNA(Gln) | -0.026208 |
| 218 | COG0779 | Required for maturation of 30S ribosomal subunits | -0.026200 |
| 219 | COG4346 | Dolichyl-phosphate-mannose--protein O-mannosyl transferase | -0.026177 |
| 220 | COG2723 | 6-phospho-beta-galactosidase activity | 0.026135 |
| 221 | COG4096 | Type I site-specific restriction-modification system, R (Restriction) subunit and related | -0.026123 |
| 222 | 33S26 | HNH nucleases | -0.026119 |
| 223 | COG0312 | modulator of DNA gyrase | 0.026070 |
| 224 | COG3710 | Transcriptional regulator | 0.026058 |
| 225 | COG4303 | Ethanolamine ammonia lyase large subunit | 0.026018 |
| 226 | COG0674 | pyruvate-flavodoxin oxidoreductase activity | -0.025983 |
| 227 | COG1072 | (Pantothenic acid kinase)) | -0.025965 |
| 228 | COG1640 | 4-alpha-glucanotransferase | 0.025960 |
| 229 | COG3394 | polysaccharide catabolic process | 0.025900 |
| 230 | COG0294 | Catalyzes the condensation of para-aminobenzoate (pABA) with 6-hydroxymethyl-7,8-dihydropterin diphosphate (DHPt-PP) to form 7,8-dihydropteroate (H2Pte), the immediate precursor of folate derivatives | 0.025874 |
| 231 | COG2819 | hydrolase of the alpha beta superfamily | 0.025840 |
| 232 | COG5018 | Exonuclease | 0.025820 |
| 233 | COG1203 | CRISPR-associated helicase, Cas3 | 0.025737 |
| 234 | COG3807 | Bacterial SH3 domain | -0.025693 |
| 235 | COG1394 | ATPase activity, coupled to movement of substances | -0.025678 |
| 236 | COG2894 | cell division | 0.025670 |
| 237 | COG0447 | Converts o-succinylbenzoyl-CoA (OSB-CoA) to 1,4- dihydroxy-2-naphthoyl-CoA (DHNA-CoA) | -0.025660 |
| 238 | COG3541 | Predicted nucleotidyltransferase | -0.025614 |
| 239 | COG5627 | protein ubiquitination | -0.025590 |
| 240 | COG2706 | 6-phosphogluconolactonase activity | -0.025534 |
| 241 | COG1869 | D-ribose catabolic process | 0.025471 |
| 242 | COG3682 | Transcriptional regulator | -0.025432 |
| 243 | COG2146 | nitrite reductase [NAD(P)H] activity | -0.025393 |
| 244 | COG2313 | pseudouridylate synthase activity | -0.025382 |
| 245 | COG0001 | glutamate-1-semialdehyde 2,1-aminomutase activity | 0.025355 |
| 246 | COG0830 | nickel cation binding | -0.025326 |
| 247 | COG0144 | Specifically methylates the cytosine at position 967 (m5C967) of 16S rRNA | -0.025321 |
| 248 | COG4744 | Conserved Protein | -0.025319 |
| 249 | COG0692 | Excises uracil residues from the DNA which can arise as a result of misincorporation of dUMP residues by DNA polymerase or due to deamination of cytosine | -0.025267 |
| 250 | 308PC | Domain of unknown function (DUF4956) | 0.025264 |